A global rise in antimicrobial resistant infections has resulted in an estimated 4.95 million deaths in 2019.1,2 The WHO 2024 priority pathogens list includes 24 notorious antimicrobial resistant organisms identified as targets for research and development (R&D) and public health interventions.2 The list aims to address the ever-changing landscape of global antimicrobial resistance (AMR) and prioritize R&D and investments in AMR.
The WHO priority pathogens list includes 3 “critical” organisms: carbapenem-resistant Acinetobacter baumannii, cephalosporin-resistant Enterobacterales, and carbapenem-resistant Enterobacterales (CRE).2 While the development of new antimicrobials is crucial to combating these difficult-to-treat pathogens, this action will not be enough to combat AMR. Infection prevention and control measures and enacting robust stewardship measures are required to ensure appropriate use of new antimicrobials and sustain their longevity in clinical practice.2
Intertwined with antimicrobial stewardship is diagnostic stewardship, which refers to ordering the right tests for the right patient at the right time to inform optimal clinical care.3 More specifically, diagnostic stewardship aims to reduce diagnostic error, which is defined as delayed, missed, or wrong diagnoses that can lead to patient harm.3 The CDC Core Elements of Hospital Antibiotic Stewardship Programs (ASPs) also identifies diagnostic stewardship as an integral component of antimicrobial stewardship.4 Investing and implementing rapid diagnostics and timely antimicrobial susceptibility testing (AST) of clinically relevant multidrug-resistant (MDR) pathogens are essential to establishing effective therapy. Novel molecular methods for pathogen-specific or syndrome-based rapid diagnostic testing (RDT) and rapid antimicrobial susceptibility testing (RAST) are described in this review.
Rapid Diagnostic Tests
While conventional culture remains the gold standard for diagnosing bacterial infection, this method is hindered by several limitations, including limited sensitivity and long turnaround times for pathogen recovery, identification, and AST, which can take up to several days, delaying time-sensitive, lifesaving treatment.1 Non–culture-based diagnostics, including RDTs, have emerged in clinical microbiology laboratories and may represent a potential solution to slow turnaround times. RDTs are defined by their performance features rather than by the testing platform or test analyte they use.5 They are characterized by their quick turnaround times, usually within 2 hours, which help guide clinical decision making and allow for immediate management at the point of care.6 RDTs come in different formats and platforms, catering to various detection targets. They are intended for identifying:
- pathogen-specific antigens;
- nucleic acid sequences (ie, molecular testing); or
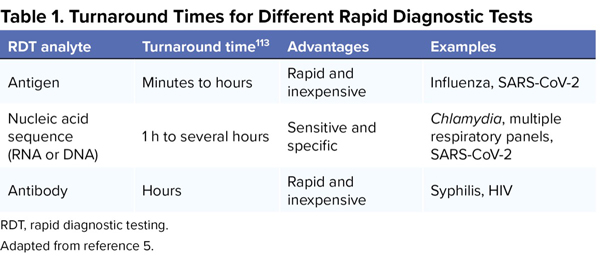
- detecting host antibodies in response to specific pathogens (Table 1).5
| Table 1. Turnaround Times for Different Rapid Diagnostic Tests | |||
| RDT analyte | Turnaround time113 | Advantages | Examples |
|---|---|---|---|
| Antigen | Minutes to hours | Rapid and inexpensive | Influenza, SARS-CoV-2 |
| Nucleic acid sequence (RNA or DNA) | 1 h to several hours | Sensitive and specific | Chlamydia, multiple respiratory panels, SARS-CoV-2 |
| Antibody | Hours | Rapid and inexpensive | Syphilis, HIV |
| RDT, rapid diagnostic testing. Adapted from reference 5. | |||
Rapid antigen tests have existed for years for upper respiratory viruses (ie, influenza and respiratory syncytial virus [RSV]) and streptococcal pharyngitis.7 RDTs for antibody and antigen detection are less sensitive than standard laboratory assays, and positive results should be confirmed with molecular testing (ie, nucleic acid amplification tests).5 Therefore, the focus of this review is on molecular testing for respiratory and bloodstream infections (BSIs) as well as novel methods for resistance detection and RAST.
Over the last decade, there has been an increase in the number of FDA-cleared molecular diagnostics for acute respiratory tract infections (RTIs).8 Specifically, the COVID-19 pandemic spurred a molecular point-of-care and laboratory testing revolution as evidenced by numerous molecular tests, including nucleic acid amplification tests, gaining emergency use authorization from the FDA.9 Molecular tests capable of detecting 1 or more respiratory viruses have become the diagnostic gold standard in clinical virology, as their results are highly sensitive and specific (Table 1).5,8 Until recently, molecular tests have been limited to viral pathogens and a few atypical bacteria including Mycoplasma pneumoniae, Chlamydia pneumoniae, and Bordetella spp. As a result, limited high-quality data exist about test performance and clinical impact of new molecular tests for bacterial pathogens. With the exception of nasal methicillin-resistant Staphylococcus aureus (MRSA) screening, current guidelines do not address molecular testing for bacterial pathogens in patients with hospital-acquired or ventilator-associated bacterial pneumonia.10,11 However, new molecular tests for respiratory bacterial pathogens, including multiplex respiratory panels, have started to accumulate high-quality data in randomized controlled trials and meta-analyses.12-14,15-20
The etiology of RTIs has been determined through traditional culture-based methods, which involves the isolation and growth of bacterial colonies on agar plates and subsequent AST. This process usually takes 24 to 48 hours for microorganism identification and another 24 to 48 hours for susceptibility testing.21 During this period, patients receive empiric antimicrobial therapy that may be suboptimal or overly broad spectrum, which can lead to the development of AMR and adverse events, such as Clostridioides difficile infection.22,23 As a result, novel molecular testing methods, such as multiplex panels, have been associated with reduced time to results and length of stay (LOS) as well as improved infection control.14 Despite the benefits of molecular testing, cost remains a barrier for implementing RDTs in the clinical setting.24
Viral pathogens, including rhinovirus, influenza, coronavirus, and RSV, are the most common cause of upper RTIs (URTIs).25 URTIs are among the top 3 diagnoses in the outpatient setting and account for an estimated 10 million outpatient visits per year in the United States.26 For this reason, healthcare professionals should be cautious about overusing or misusing antibiotics because fewer than 10% of URTIs are attributed to a bacterial cause.27,28 Inappropriate antibiotic prescribing is still common, reaching more than 40% in outpatients with acute URTIs.29
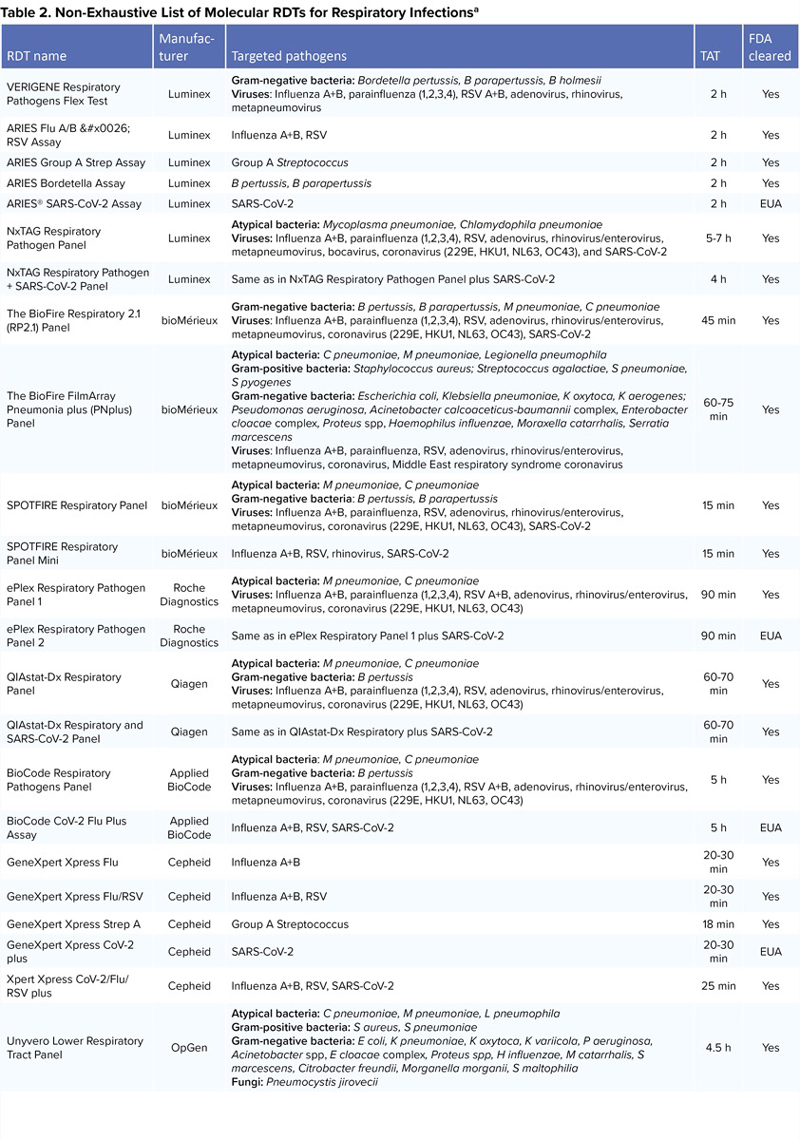
Rapid multiplex panels allow for the accurate detection of 10 or more respiratory pathogens simultaneously. The multiplex panel may also incorporate a “sample-to-answer” design, whereby extraction, amplification, and analysis of specimens are integrated fully within closed processes.14 These sample-to-answer systems allow for the accurate detection of multiple respiratory targets with similar presenting symptoms in less than 1 hour, expediting earlier discharge and appropriate treatment.30-32 Examples of syndromic multiplex panels capable of detecting 10 or more common URTI pathogens include the VERIGENE Respiratory Pathogens Flex Test (Luminex), NxTAG Respiratory Pathogen Panel (Luminex), BioFire Respiratory 2.1 (RP2.1) Panel (bioMérieux), SPOTFIRE Respiratory Panel (bioMérieux), ePlex Respiratory Pathogen Panel (Roche Diagnostics), QIAstat-Dx Respiratory Panel (Qiagen), and BioCode Respiratory Pathogens Panel (Applied BioCode) (Table 2). These RDTs can detect common viral pathogens associated with URTIs such as influenza A, influenza B, parainfluenza 1, parainfluenza 2, parainfluenza 3, parainfluenza 4, RSV A and B, adenovirus, rhinovirus, and metapneumovirus. However, they differ in the technology used, result turnaround time, and their ability to detect additional pathogens.33-37 For example, the ePlex Respiratory Pathogen Panel and NxTAG Respiratory Pathogen Panel cannot identify Bordetella spp.38,39 Moreover, the ePlex Respiratory Pathogen Panel cannot identify atypical bacteria (M pneumoniae, C pneumoniae), while the NxTAG Respiratory Pathogen Panel is the only product listed that can identify bocavirus.40-43 The BioFire RP2.1 Panel was the first FDA-cleared panel that could identify SARS-CoV-2. The RDTs also differ from each other regarding the speed of sample processing and result reporting. The result turnaround time can range from 15 minutes to 5 to 7 hours. Refer to Table 2 for a non-exhaustive list and comparison of common RDTs designed for RTIs.
| Table 2. Non-Exhaustive List of Molecular RDTs for Respiratory Infectionsa | ||||
| RDT name | Manufacturer | Targeted pathogens | TAT | FDA cleared |
|---|---|---|---|---|
| VERIGENE Respiratory Pathogens Flex Test | Luminex | Gram-negative bacteria: Bordetella pertussis, B parapertussis, B holmesii Viruses: Influenza A+B, parainfluenza (1,2,3,4), RSV A+B, adenovirus, rhinovirus, metapneumovirus | 2 h | Yes |
| ARIES Flu A/B & RSV Assay | Luminex | Influenza A+B, RSV | 2 h | Yes |
| ARIES Group A Strep Assay | Luminex | Group A Streptococcus | 2 h | Yes |
| ARIES Bordetella Assay | Luminex | B pertussis, B parapertussis | 2 h | Yes |
| ARIES® SARS-CoV-2 Assay | Luminex | SARS-CoV-2 | 2 h | EUA |
| NxTAG Respiratory Pathogen Panel | Luminex | Atypical bacteria: Mycoplasma pneumoniae, Chlamydophila pneumoniae Viruses: Influenza A+B, parainfluenza (1,2,3,4), RSV, adenovirus, rhinovirus/enterovirus, metapneumovirus, bocavirus, coronavirus (229E, HKU1, NL63, OC43), and SARS-CoV-2 | 5-7 h | Yes |
| NxTAG Respiratory Pathogen + SARS-CoV-2 Panel | Luminex | Same as in NxTAG Respiratory Pathogen Panel plus SARS-CoV-2 | 4 h | Yes |
| The BioFire Respiratory 2.1 (RP2.1) Panel | bioMérieux | Gram-negative bacteria: B pertussis, B parapertussis, M pneumoniae, C pneumoniae Viruses: Influenza A+B, parainfluenza (1,2,3,4), RSV, adenovirus, rhinovirus/enterovirus, metapneumovirus, coronavirus (229E, HKU1, NL63, OC43), SARS-CoV-2 | 45 min | Yes |
| The BioFire FilmArray Pneumonia plus (PNplus) Panel | bioMérieux | Atypical bacteria: C pneumoniae, M pneumoniae, Legionella pneumophila Gram-positive bacteria: Staphylococcus aureus; Streptococcus agalactiae, S pneumoniae, S pyogenes Gram-negative bacteria: Escherichia coli, Klebsiella pneumoniae, K oxytoca, K aerogenes; Pseudomonas aeruginosa, Acinetobacter calcoaceticus-baumannii complex, Enterobacter cloacae complex, Proteus spp, Haemophilus influenzae, Moraxella catarrhalis, Serratia marcescens Viruses: Influenza A+B, parainfluenza, RSV, adenovirus, rhinovirus/enterovirus, metapneumovirus, coronavirus, Middle East respiratory syndrome coronavirus | 60-75 min | Yes |
| SPOTFIRE Respiratory Panel | bioMérieux | Atypical bacteria: M pneumoniae, C pneumoniae Gram-negative bacteria: B pertussis, B parapertussis Viruses: Influenza A+B, parainfluenza, RSV, adenovirus, rhinovirus/enterovirus, metapneumovirus, coronavirus (229E, HKU1, NL63, OC43), SARS-CoV-2 | 15 min | Yes |
| SPOTFIRE Respiratory Panel Mini | bioMérieux | Influenza A+B, RSV, rhinovirus, SARS-CoV-2 | 15 min | Yes |
| ePlex Respiratory Pathogen Panel 1 | Roche Diagnostics | Atypical bacteria: M pneumoniae, C pneumoniae Viruses: Influenza A+B, parainfluenza (1,2,3,4), RSV A+B, adenovirus, rhinovirus/enterovirus, metapneumovirus, coronavirus (229E, HKU1, NL63, OC43) | 90 min | Yes |
| ePlex Respiratory Pathogen Panel 2 | Roche Diagnostics | Same as in ePlex Respiratory Panel 1 plus SARS-CoV-2 | 90 min | EUA |
| QIAstat-Dx Respiratory Panel | Qiagen | Atypical bacteria: M pneumoniae, C pneumoniae Gram-negative bacteria: B pertussis Viruses: Influenza A+B, parainfluenza (1,2,3,4), RSV, adenovirus, rhinovirus/enterovirus, metapneumovirus, coronavirus (229E, HKU1, NL63, OC43) | 60-70 min | Yes |
| QIAstat-Dx Respiratory and SARS-CoV-2 Panel | Qiagen | Same as in QIAstat-Dx Respiratory plus SARS-CoV-2 | 60-70 min | Yes |
| BioCode Respiratory Pathogens Panel | Applied BioCode | Atypical bacteria: M pneumoniae, C pneumoniae Gram-negative bacteria: B pertussis Viruses: Influenza A+B, parainfluenza (1,2,3,4), RSV A+B, adenovirus, rhinovirus/enterovirus, metapneumovirus, coronavirus (229E, HKU1, NL63, OC43) | 5 h | Yes |
| BioCode CoV-2 Flu Plus Assay | Applied BioCode | Influenza A+B, RSV, SARS-CoV-2 | 5 h | EUA |
| GeneXpert Xpress Flu | Cepheid | Influenza A+B | 20-30 min | Yes |
| GeneXpert Xpress Flu/RSV | Cepheid | Influenza A+B, RSV | 20-30 min | Yes |
| GeneXpert Xpress Strep A | Cepheid | Group A Streptococcus | 18 min | Yes |
| GeneXpert Xpress CoV-2 plus | Cepheid | SARS-CoV-2 | 20-30 min | EUA |
| Xpert Xpress CoV-2/Flu/RSV plus | Cepheid | Influenza A+B, RSV, SARS-CoV-2 | 25 min | Yes |
| Unyvero Lower Respiratory Tract Panel | OpGen | Atypical bacteria: C pneumoniae, M pneumoniae, L pneumophila Gram-positive bacteria: S aureus, S pneumoniae Gram-negative bacteria: E coli, K pneumoniae, K oxytoca, K variicola, P aeruginosa, Acinetobacter spp, E cloacae complex, Proteus spp, H influenzae, M catarrhalis, S marcescens, Citrobacter freundii, Morganella morganii, S maltophilia Fungi: Pneumocystis jirovecii | 4.5 h | Yes |
| EUA, emergency use authorization; MRSA, methicillin-resistant Staphylococcus aureus; RDT, rapid diagnostic testing; RSV, respiratory syncytial virus; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; TAT, turnaround time. a FDA cleared or authorized under an EUA. | ||||
Despite the variation among these products regarding the turnaround time, all can provide results hours to days faster than traditional diagnostic methods. In addition, these rapid testing panels show high sensitivity and specificity compared with traditional laboratory-developed real-time polymerase chain reaction (PCR) assays. For example, the BioFire RP2.1 Panel showed a sensitivity of 97.1% and specificity of 99.3%, while the NxTAG Respiratory Pathogen Panel showed a sensitivity of 95.2% and specificity of 99.6%.35,44 A retrospective study by Nijhuis et al compared the ePlex Respiratory Pathogen Panel with laboratory-developed real-time PCR assays for the detection of 464 respiratory pathogens from 323 positive clinical specimens.45 Results showed that the ePlex panel had 97.4% agreement with the real-time PCR assay. Furthermore, the panel detected 17 more pathogens than the laboratory-developed real-time PCR assays.45 Another study compared the performance of the ePlex Respiratory Pathogen Panel with the BioFire RP2.1 for the detection of viral and bacterial respiratory tract pathogens in nasopharyngeal swabs. The results demonstrated overall agreement between both panels (>95%) for all targets. The overall positive percent agreement (PPA) of the ePlex Respiratory Pathogen Panel compared with the BioFire RP2.1 for viruses ranged from 85.1% (95% CI, 80.2%-88.9%) to 95.1% (95% CI, 89.0%-97.9%), while the negative percent agreement (NPA) ranged from 99.5% (95% CI, 99.1%-99.7%) to 99.8% (95% CI, 99.5%-99.9%).46
All panels mentioned above are FDA cleared for use with nasopharyngeal swab specimens. They are considered qualitative molecular tests that use multiplex nucleic acid or PCR technology, except for the BioCode Respiratory Pathogens Panel, which uses a bioinformatics approach involving DNA or RNA sequencing techniques. Ultimately, multiplex panels offer great promise in the optimization of the diagnosis and treatment of URTIs likely leading to further downstream benefits such as improved antimicrobial stewardship and patient outcomes.
In the United States, pneumonia has been a major contributor to morbidity and mortality, causing about 63,000 deaths and 1.2 million hospitalizations yearly.47,48 The American Thoracic Society/Infectious Diseases Society of America (ATS/IDSA) recommendations for empiric antimicrobial therapy in community-acquired pneumonia (CAP) are based on selecting agents targeted against the major treatable respiratory bacterial pathogens.11 While bacteria are implicated most commonly in adults with CAP, roughly one-third of cases are due to viruses.49 Unfortunately, overuse of antibiotics is common in lower RTIs, because it is difficult to distinguish between bacterial and viral etiologies due to similar manifestations.50 Antibiotic therapy may be safely withheld in patients with isolated viral pneumonia if these infections can be easily identified from those with concomitant bacterial etiology.51
Multiple syndromic molecular testing panels for respiratory bacterial pathogens are available, including the BioFire FilmArray Pneumonia Panel (bioMérieux) and Unyvero Lower Respiratory Tract Panel (OpGen), which offer increased sensitivity over traditional cultures and provide the presence of resistance markers within as little as 1 to 5 hours from specimen collection and testing. The BioFire panel offers detection of 8 viruses, 8 resistance genes, 3 atypical bacteria using qualitative targets, and 15 bacterial targets with semiquantitative analysis that can help in evaluating colonization versus infection. The Unyvero panel includes the detection of 29 bacterial pathogens and 19 resistance genes. Both panels can be used with multiple specimen types. Although semiquantitative analysis may improve the clinical specificity, neither molecular testing panels nor culture separate airway colonizers from invasive pathogens. Therefore, the possibility of a downstream effect of paradoxically increasing antimicrobial use should be a notable concern. These panels may be most useful where patients have new or worsening lung infiltrates, are moderately to severely ill, have received empiric antibiotics before obtaining cultures, and/or there is concern for multidrug-resistant bacteria or a polymicrobial infection.52
The BioFire FilmArray Pneumonia Panel showed a PPA and NPA of 98.1% and 96.2%, respectively, for identifying bacterial targets on bronchoalveolar lavage specimens compared with culture.53 Similarly, a high overall agreement of 99.2% (95% CI, 98.4%-99.6%) for viral detection was seen between the FilmArray panel and culture.54 The excellent sensitivity of the pneumonia panel may help rule out bacterial coinfections and avoid inappropriate antibiotic prescribing, but positive results should be interpreted with caution. In a multinational study of 52 laboratories, the panel identified significantly more positive specimens (76.1%) than standard-of-care (SOC) testing (56.03%; P=0.0001) and more potential pathogens than SOC (P=0.0001) independent of specimen type with the most significant discrepancies for fastidious pathogens.55 Lower SOC bacteria detection may be explained by local reporting guidelines and testing of specimens from patients on antibiotics.55 The FilmArray Pneumonia Panel carries interpretation challenges, including understanding the increased detection rates, colonization, and infection differentiation.55 A recent study prospectively examined sputum specimens in 70 patients with pneumonia, and the potential for antibiotic optimization was observed in 56 of 70 patients.56 Nine bacteria in 9 patients were not covered by empiric therapy, and 70 antibiotics in 49 patients could have been discontinued.56 Likewise, a retrospective multicenter study observed antimicrobial de-escalation in 63 of 159 (40%) and escalation in 35 (22%) hospitalized pneumonia patients based on FilmArray panel results.57 These results reinforce the potential of the panel to reduce unnecessary antimicrobial exposure and increase the appropriateness of empiric antibiotic therapy.
The Unyvero panel also has reported robust diagnostic accuracy. In an evaluation by Collins et al, the PPA and NPA for the bacterial targets was 96.5% and 99.6%, respectively.58 Also, Klein et al found a high overall PPA and NPA with culture, but 21.7% of specimens had additional potential pathogens identified by the panel.59 Moreover, the Unyvero panel facilitated earlier initiation of effective antibiotic therapy in 20 of 95 patients (21%) and de-escalation in 37 patients (39%) with ventilator-associated pneumonia in a prospective study.60 In another study by Pickens et al, the Unyvero panel predicted antibiotic de-escalation in 65.9% of patients (405/615), where unnecessary MRSA coverage (69%) and Pseudomonas aeruginosa coverage (64%) were prevalent.61
Several new RDT panels are being developed to detect respiratory pathogens in lower RTIs, including the AusDiagnostics Pneumonia 16-well panel and the comprehensive Respiratory Pathogens 24-well panel, which uses novel multiplex tandem PCR technology.62 Challenges with the interpretation of newer RDTs are of concern but may be mitigated by ASP intervention.
RDTs for BSIs
BSIs refer to bacteremia or fungemia in the bloodstream. The blood is considered a sterile environment, and BSIs can result from infections in other parts of the body, catheter use, contaminated surgical procedures, or simply through breaks in the skin.63,64 BSIs are serious conditions associated with high morbidity and mortality globally.65,66 Blood culture remains the gold standard for BSI diagnosis, but it has limitations such as slow turnaround time and reduced sensitivity. Some studies estimate the sensitivity of blood culture to be as low as 50%.67,68 RDTs for BSIs can improve the sensitivity of pathogen detection and shorten result reporting time significantly.69 A recently published series of network meta-analyses included 88 studies of patients with BSIs with the aim of comparing the clinical impact of RDTs to conventional blood culture in association with ASPs.70 A total of 25,682 patient encounters were included and matrix-assisted laser desporption/ionization time-of-flight (MALDI-TOF) mass spectrometry was considered an RDT when applied directly on a positive blood culture. The ASP was defined as any intervention aimed at implementing the RDT or blood culture results, by means of recommendations from the ASP team. VERIGENE, Xpert, Accelerate, and BCID (Blood Culture Identification) were among the commonly used RDTs in this study.70 The results showed a significant reduction in mortality associated with RDT versus blood culture, even when both are embedded within an ASP (odds ratio, 0.78; 95% CI, 0.63-0.96). Prior to this study, the relative contribution of RDTs and ASPs on improved survival was unknown. With these results, it may be beneficial to adopt RDTs and ASPs even in settings with an established ASP in association with conventional blood culture. However, the results should be extrapolated carefully, as the positive impact of RDTs may be specific to the setting or type of RDT or ASP only.70
Molecular RDTs have transformed the approach to managing BSIs and address potential contaminants in blood cultures, such as coagulase-negative staphylococci. RDTs can provide crucial, actionable insights at significantly earlier stages of treatment when compared with traditional microbiological culture methods. The implementation of molecular RDTs has been associated with decreases in time to effective therapy, hospital LOS, and mortality when accompanied by ASP interventions.71 Similar to the clinical impacts observed, a cost-effectiveness analysis also has reflected the benefits of molecular RDTs in BSIs.72 These data also highlight that the strong synergy of stewardship and RDTs has an 80% chance of cost-effectiveness with an ASP, but only 41.1% without. Alternatively, automated rapid phenotypic testing systems such as the Accelerate PhenoTest system can yield organism identification, minimum inhibitory concentration (MIC), and susceptibility interpretation with a turnaround time of approximately 7 hours after positive blood culture.73
RDTs developed for BSIs use different diagnostic technologies, including PCR-based technologies (eg, BioFire FilmArray BCID, the Roche Diagnostic cobas ePlex BCID), Multiplex Tandem PCR (eg, AusDiagnostics), nanoparticle probe technology (eg, VERIGENE BC-GP and BC-GN [gram-positive and gram-negative blood culture nucleic acid tests]), MALDI-TOF (eg,, bioMérieux, BD Bruker), magnetic resonance technology (eg, T2Candida panel and T2Bacteria panel), or morphokinetic cellular analysis technique (eg, Accelerate PhenoTest BC Kit). Although all of these RDTs can process specimens rapidly and provide results in just a few hours from positive blood culture, T2 panels (T2Candida and T2Bacteria) are the only FDA-approved diagnostic panels that can process samples directly from whole blood without waiting for positive blood culture, substantially expediting result reporting.
This can be important in patients where rapid administration of directed therapy is crucial due to severe infection (eg, septic shock).74 However, T2 panels can only identify a small number of pathogens, 6 bacteria and 5 fungi, compared with larger panels like the ePlex panel, which detects 20 gram-positive and 21 gram-negative bacterial pathogens. Among those RDTs that can only process positive blood culture samples, the largest difference in pathogen detection time is 1.5 hours. The BioFire BCID2 panel, which can detect 15 gram-negative bacteria, 11 gram-positive bacteria, 7 yeasts with the fastest turnaround time (1 hour), showed improved outcomes in identifying gram-negative bacteria causing BSIs when compared with the VERIGENE panel (detects 9 gram-negative bacteria, 12 gram-positive bacteria) and the BioFire original BCID panel (detects 11 gram-negative bacteria, 8 gram-positive bacteria, 5 yeasts).75
Some RDTs may include pathogens in their detection list but are not FDA cleared for that specific pathogen identification. For example, the VERIGENE Bloodstream Infections Testing Panel is not FDA cleared for Serratia marcescens or Micrococcus spp. Furthermore, some tests may not be able to distinguish among certain bacteria. For example, the VERIGENE Bloodstream Infections Testing Panel cannot distinguish Escherichia coli from Shigella spp.
Of note, only the BioFire BCID2 panel and ePlex BCID Panels can identify Stenotrophomonas maltophilia. Similarly, for Candida auris, only the BioFire BCID2 panel, ePlex BCID Panels, and MALDI Biotyper MT Sepsityper Kit can identify this notorious organism. Refer to Table 3 for a direct comparison among common RDTs for BSIs. New RDTs are in development to further reduce time to results to less than 1 hour. For example, the FAST-ID BSI Panel (Qvella) can identify more than 90% of pathogens causing BSIs directly from a whole blood sample. Likely, the use of non–culture-based methods in RDT will lead to decreased time to appropriate therapy and may have a positive effect on health outcomes, specifically in critically ill patients.
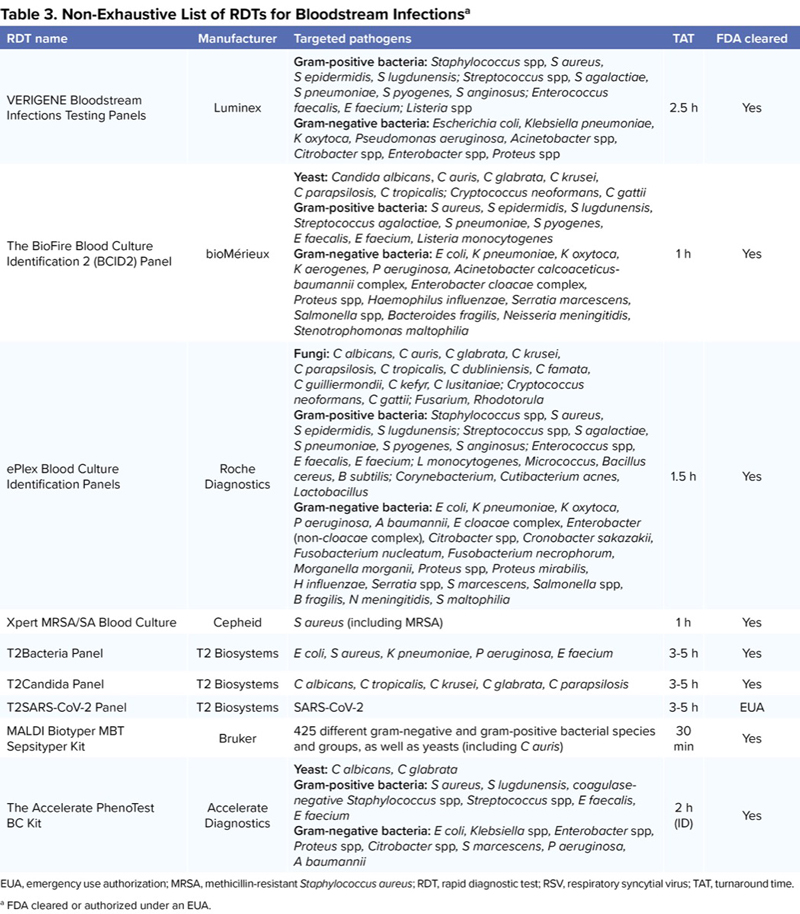
| Table 3. Non-Exhaustive List of RDTs for Bloodstream Infectionsa | ||||
| RDT name | Manufacturer | Targeted pathogens | TAT | FDA cleared |
|---|---|---|---|---|
| VERIGENE Bloodstream Infections Testing Panels | Luminex | Gram-positive bacteria: Staphylococcus spp, S aureus, S epidermidis, S lugdunensis; Streptococcus spp, S agalactiae, S pneumoniae, S pyogenes, S anginosus; Enterococcus faecalis, E faecium; Listeria spp Gram-negative bacteria: Escherichia coli, Klebsiella pneumoniae, K oxytoca, Pseudomonas aeruginosa, Acinetobacter spp, Citrobacter spp, Enterobacter spp, Proteus spp | 2.5 h | Yes |
| The BioFire Blood Culture Identification 2 (BCID2) Panel | bioMérieux | Yeast: Candida albicans, C auris, C glabrata, C krusei, C parapsilosis, C tropicalis; Cryptococcus neoformans, C gattii Gram-positive bacteria: S aureus, S epidermidis, S lugdunensis, Streptococcus agalactiae, S pneumoniae, S pyogenes, E faecalis, E faecium, Listeria monocytogenes Gram-negative bacteria: E coli, K pneumoniae, K oxytoca, K aerogenes, P aeruginosa, Acinetobacter calcoaceticus-baumannii complex, Enterobacter cloacae complex, Proteus spp, Haemophilus influenzae, Serratia marcescens, Salmonella spp, Bacteroides fragilis, Neisseria meningitidis, Stenotrophomonas maltophilia | 1 h | Yes |
| ePlex Blood Culture Identification Panels | Roche Diagnostics | Fungi: C albicans, C auris, C glabrata, C krusei, C parapsilosis, C tropicalis, C dubliniensis, C famata, C guilliermondii, C kefyr, C lusitaniae; Cryptococcus neoformans, C gattii; Fusarium, Rhodotorula Gram-positive bacteria: Staphylococcus spp, S aureus, S epidermidis, S lugdunensis; Streptococcus spp, S agalactiae, S pneumoniae, S pyogenes, S anginosus; Enterococcus spp, E faecalis, E faecium; L monocytogenes, Micrococcus, Bacillus cereus, B subtilis; Corynebacterium, Cutibacterium acnes, Lactobacillus Gram-negative bacteria: E coli, K pneumoniae, K oxytoca, P aeruginosa, A baumannii, E cloacae complex, Enterobacter (non-cloacae complex), Citrobacter spp, Cronobacter sakazakii, Fusobacterium nucleatum, Fusobacterium necrophorum, Morganella morganii, Proteus spp, Proteus mirabilis, H influenzae, Serratia spp, S marcescens, Salmonella spp, B fragilis, N meningitidis, S maltophilia | 1.5 h | Yes |
| Xpert MRSA/SA Blood Culture | Cepheid | S aureus (including MRSA) | 1 h | Yes |
| T2Bacteria Panel | T2 Biosystems | E coli, S aureus, K pneumoniae, P aeruginosa, E faecium | 3-5 h | Yes |
| T2Candida Panel | T2 Biosystems | C albicans, C tropicalis, C krusei, C glabrata, C parapsilosis | 3-5 h | Yes |
| T2SARS-CoV-2 Panel | T2 Biosystems | SARS-CoV-2 | 3-5 h | EUA |
| MALDI Biotyper MBT Sepsityper Kit | Bruker | 425 different gram-negative and gram-positive bacterial species and groups, as well as yeasts (including C auris) | 30 min | Yes |
| The Accelerate PhenoTest BC Kit | Accelerate Diagnostics | Yeast: C albicans, C glabrata Gram-positive bacteria: S aureus, S lugdunensis, coagulase-negative Staphylococcus spp, Streptococcus spp, E faecalis, E faecium Gram-negative bacteria: E coli, Klebsiella spp, Enterobacter spp, Proteus spp, Citrobacter spp, S marcescens, P aeruginosa, A baumannii | 2 h (ID) | Yes |
| EUA, emergency use authorization; MRSA, methicillin-resistant Staphylococcus aureus; RDT, rapid diagnostic test; RSV, respiratory syncytial virus; TAT, turnaround time. a FDA cleared or authorized under an EUA. | ||||
Moreover, in addition to using RDTs for detection and early initiation of the appropriate antimicrobial agents, they can be used for early discontinuation of empiric therapy. For example, the use of the T2Candida panel resulted in a reduction in empiric antifungal days of therapy in ICU patients compared with using the beta-D-glucan biomarker.76 However, RDTs that can decrease the turnaround time for biomarker detection (eg, beta-D-glucan, galactomannan) could also help to rule out infection and facilitate the early discontinuation of empiric antimicrobial therapy. For example, Fungitell STAT is an FDA-cleared assay developed by ACC, which provides rapid screening for invasive candidiasis through the detection of serum beta-D-glucan in about 1 hour. This test can help detect intrabdominal candidiasis in ICU patients in whom blood culture sensitivity is low, and beta-D-glucan testing has demonstrated utility.77,78
A retrospective study by Walker et al evaluated the clinical impact of VERIGENE BC-GN panel implementation for the detection of gram-negative bacteria in positive blood cultures in hospitalized patients by comparing the 6-month period before and the 6-month period after implementing the panel and demonstrated that the BC-GN panel identified 87% of gram-negative cultures and was accurate in 98% of these cultures compared with a conventional culture method. The BC-GN panel was associated with faster pathogen identification than the traditional culture method (10.9 vs 37.9 hours; P<0.001). Furthermore, LOS and 30-day mortality were significantly lower after implementing the BC-GN panel (P<0.05).79 Another retrospective study compared the turnaround time and accuracy of organism and resistance gene detection for VERIGENE Bloodstream Infections Testing Panels and the BioFire Blood Culture Identification (original panel) with the traditional culture method for 173 patients with positive blood cultures. This study found that both panels accurately identified pathogens much faster than the traditional culture method by 27.95 and 29.17 hours for the VERIGENE and BioFire panels, respectively.80 One study compared 3 RDT panels (VERIGENE BC-GN, BioFire BCID, and BioFire BCID2) using the Desirability of Outcome Ranking Management of Antimicrobial Therapy (DOOR MAT) to assess potential downstream antimicrobial prescribing decisions resulting from the panel’s different organism and resistance detection. They found that the mean DOOR MAT score was higher, where a higher score reflects better outcome, for BioFire BCID2 than VERIGENE BC-GN (P<0.7) and BioFire BCID (P<0.0001). The mean DOOR MAT score was 83.8 (SD, 25.7), 59.9 (SD, 33.7), and 89.7 (SD, 24.7) for VERIGENE BC-GN, BioFire BCID, and BioFire BCID2, respectively.75
A study by Huang et al evaluated the ePlex BCID Panels for the detection of BSI pathogens compared with traditional culture for 216 patients with positive blood culture. This study found that the PPA was 96% and 94% for gram-positive and gram-negative bacteria, respectively.81 Another recent study evaluated the turnaround time of the ePlex BCID-GN panel compared with MALDI-TOF for gram-negative bacteria identification direct from blood culture. The study showed that the ePlex BCID-GN panel identified gram-negative bacteria 26.7 hours earlier on average compared with MALDI-TOF.82 However, the ePlex panel, along with the other aforementioned multiplex panels, require an additional 12 to 48 hours of incubation time for blood culture prior to providing results.83 Alternatively, the turnaround time can be further reduced by using direct from whole blood samples, such as those samples used by T2Biosystems, eliminating the time associated with blood culture. A recent prospective multicenter study compared the T2Candida panel with beta-D-glucan for diagnosis of intraabdominal candidiasis in critically ill patients using blood samples. The results demonstrated that T2Candida performance was comparable to beta-D-glucan for intraabdominal candidiasis diagnosis. Of the 134 patients evaluated, 13 had proven intraabdominal candidiasis, while 2 had concurrent candidemia. The sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV), respectively, were 46%, 97%, 61%, and 94% for T2Candida and 85%, 83%, 36%, and 98% for beta-D-glucan. All positive T2Candida results were consistent with the culture results at the species level, except for one case of dual infection.84 Furthermore, one study compared the turnaround time, sensitivity, and specificity of the T2Bacteria panel with the standard blood culture method in patients with BSIs. This study found that the mean time to species identification with T2Bacteria was 3.61 to 7.70 hours, depending on the number of samples tested, compared with the mean time from the start of blood culture incubation to positivity and species identification, which were 38.5 hours (SD, 32.8) and 71.7 hours (SD, 39.3), respectively, for the traditional method.85 Another component of BSI diagnosis and treatment is AST, and a unique feature of the Accelerate PhenoTest system is that it can provide both organism identification within 2 hours and AST results within 7 hours from positive blood culture. Several studies have explored the Accelerate PhenoTest system coupled with ASP intervention. Results of 448 patients with gram-negative BSIs in a randomized controlled trial with prospective audit and feedback in both arms reflected significantly faster antibiotic changes (median decrease of approximately 25 hours for gram-negative antibiotics; P<0.001) with the Accelerate PhenoTest system compared with culture. Antibiotic escalation was also significantly faster with the Accelerate PhenoTest system relative to culture-based methods for antimicrobial resistant BSIs (median decrease of approximately 43 hours; P=0.01). There were no differences between arms in patient outcomes, including LOS and mortality.86 These findings suggest that the Accelerate PhenoTest system may play a major role in critically ill patients who have multiple risk factors for antimicrobial drug resistance in which a timely decision to escalate therapy should be made. Likely, RDTs incorporating both organism identification as well as AST or resistance detection will further expedite appropriate treatment of BSIs.
RDTs for Resistance Detection
Bacteria can develop resistance against antimicrobial agents through various mechanisms, including enzymatic inactivation via carbapenemases (eg, KPC, NDM, IMP, VIM, OXA), altering membrane permeability (eg, porin loss), efflux pump overexpression, target site modification, and bypass of inhibition (ie, bacteria avoid using the target enzyme).87,88 Antimicrobial resistance is associated with increased healthcare costs, morbidity, and mortality.89 Knowledge of which carbapenemase is being produced can help direct treatment decisions as newer specific beta-lactam antibiotics have activity against specific carbapenemases. As a result, recent guidance by IDSA about the treatment of antimicrobial resistant gram-negative infections strongly encourages the implementation of molecular tests to identify the presence of carbapenemases in clinical CRE isolates.90 Comparative studies evaluating the different RDTs for carbapenemase detection are scarce.
One method comparison study evaluated the performance of NG-Test CARBA 5 (Hardy Diagnostics) with a composite reference method, including the Xpert CARBA-R (Cepheid), for detection of carbapenemases in 309 bacterial isolates. The results showed 100% agreement between the NG-Test CARBA 5 and the composite reference method on commonly used agar media.91 A study by Dortet et al compared the carbapenemase-producing Enterobacteriaceae rapid detection performance of 3 MALDI-TOF-based techniques (MBT STAR-Carba IVD kit [Bruker], Microflex LT Biotyper [Bruker], and VITEK MS Plus [bioMérieux]) with RAPIDEC CARBA NP. A total of 175 isolates were tested, and MALDI-TOF-based techniques demonstrated sensitivities ranging from 95% to 100% and specificities from 98.2% to 100% compared with 99.2% and 100%, respectively, for the RAPIDEC CARBA NP (bioMérieux).92 Yoo et al accurately detected resistance genes using the BioFire pneumonia panel, showing concordant results for the resistant organisms identified by culture. However, the genetic marker of antimicrobial resistance, particularly the CTX-M and carbapenemase genes, could not be definitively linked to the microorganisms detected.93 Furthermore, the surveillance of gram-negative bacterial resistance using rectal swab testing may become standard in some settings, although the clinical utility of these tests in directing therapy has yet to show significant promise.94 However, Streck ARM-D resistance detection kits are examples of RDTs that can play a beneficial role for gram-negative resistance surveillance. These kits can detect multiple beta-lactamase–associated resistance such as AmpC, OXA, CTX-M, NIM, IMP, TEM, SHV, and others.95,96
In addition to resistance detection in gram-negative infections, the early detection of MRSA can improve patient outcomes and is essential for managing gram-positive infections. For example, MRSA nasal screen methods have evolved beyond use for infection prevention and control practice to have clinical utility for routine use in de-escalations of MRSA therapy, predominantly in patients with suspected or confirmed pneumonia. Robust evidence has reflected over 95% NPV for using the nasal PCR test to rule out MRSA pneumonia.97 Therefore, the ATS/IDSA CAP guidelines endorsed the routine use of MRSA nasal PCR screening to de-escalate MRSA coverage.11 ASP implementation of this approach has been associated with a median decrease of 2.1 days of vancomycin (P<0.01).98
Other implementation results of nasal screening in suspected or confirmed pneumonia among ICU patients have been associated with $108 per patient in cost avoidance based on the cost of surveillance testing, vancomycin, and vancomycin therapeutic drug monitoring levels.99 Reviews of implementation considerations suggest fidelity of the nasal testing for 7 days after results and lack of impact of vancomycin exposure in affecting testing results.100,101 Systematic reviews and meta-analyses have also supported the use of the screen for NPV beyond pneumonia, such as in skin and soft tissue infections.102 A national study from the Veterans Affairs system has supported this concept in the largest cohort, including clinical cultures (N=561,325).103 These data showed a high overall NPV for all infection types (96.5%) and among specific infections, including BSIs (96.5%), intraabdominal infections (98.6%), respiratory infections (96.1%), wound cultures (93.1%), and urinary tract infections (99.2%). A recent study also suggests that MRSA screening can be reliably used for de-escalation within 28 days of testing, which may lead to improved diagnostic stewardship of MRSA nares PCR screening in the future.104 Similarly, Page et al evaluated the impact of the Xpert MRSA/SA blood culture test on the management of 22 positive blood cultures in obstetric patients. The study compared patients’ outcomes before and after Xpert MRSA/SA was implemented where results demonstrated a reduction in the median antimicrobial duration (55.5-43.5 hours; P=0.46) and LOS (66.5-56.0 hours; P=0.15) after RDT implementation.105 These findings suggest that RDTs for resistance detection can help decrease the frequent use of nephrotoxic antimicrobial agents like vancomycin, which is commonly used for empiric therapy for patients with septic shock.106 Table 4 is a summary and comparison of selected common RDTs designed for resistance detection.
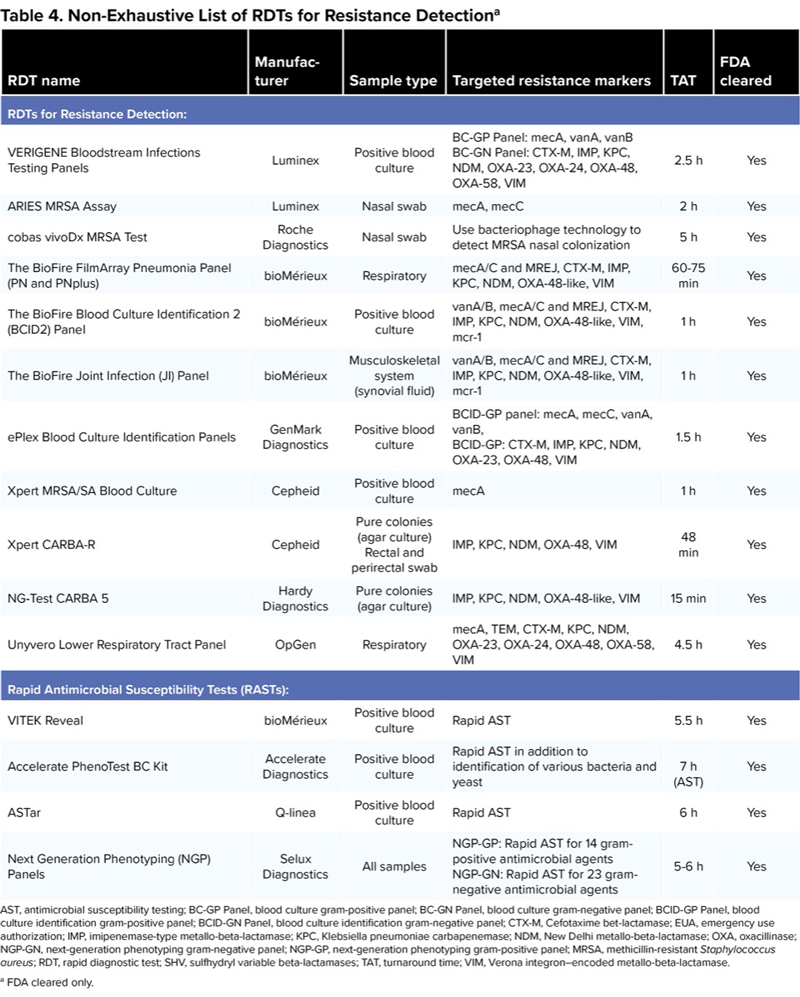
| Table 4. Non-Exhaustive List of RDTs for Resistance Detectiona | |||||
| RDT name | Manufacturer | Sample type | Targeted resistance markers | TAT | FDA cleared |
|---|---|---|---|---|---|
| RDTs for Resistance Detection: | |||||
| VERIGENE Bloodstream Infections Testing Panels | Luminex | Positive blood culture | BC-GP Panel: mecA, vanA, vanB BC-GN Panel: CTX-M, IMP, KPC, NDM, OXA-23, OXA-24, OXA-48, OXA-58, VIM | 2.5 h | Yes |
| ARIES MRSA Assay | Luminex | Nasal swab | mecA, mecC | 2 h | Yes |
| cobas vivoDx MRSA Test | Roche Diagnostics | Nasal swab | Use bacteriophage technology to detect MRSA nasal colonization | 5 h | Yes |
| The BioFire FilmArray Pneumonia Panel (PN and PNplus) | bioMérieux | Respiratory | mecA/C and MREJ, CTX-M, IMP, KPC, NDM, OXA-48-like, VIM | 60-75 min | Yes |
| The BioFire Blood Culture Identification 2 (BCID2) Panel | bioMérieux | Positive blood culture | vanA/B, mecA/C and MREJ, CTX-M, IMP, KPC, NDM, OXA-48-like, VIM, mcr-1 | 1 h | Yes |
| The BioFire Joint Infection (JI) Panel | bioMérieux | Musculoskeletal system (synovial fluid) | vanA/B, mecA/C and MREJ, CTX-M, IMP, KPC, NDM, OXA-48-like, VIM, mcr-1 | 1 h | Yes |
| ePlex Blood Culture Identification Panels | GenMark Diagnostics | Positive blood culture | BCID-GP panel: mecA, mecC, vanA, vanB, BCID-GP: CTX-M, IMP, KPC, NDM, OXA-23, OXA-48, VIM | 1.5 h | Yes |
| Xpert MRSA/SA Blood Culture | Cepheid | Positive blood culture | mecA | 1 h | Yes |
| Xpert CARBA-R | Cepheid | Pure colonies (agar culture) Rectal and perirectal swab | IMP, KPC, NDM, OXA-48, VIM | 48 min | Yes |
| NG-Test CARBA 5 | Hardy Diagnostics | Pure colonies (agar culture) | IMP, KPC, NDM, OXA-48-like, VIM | 15 min | Yes |
| Unyvero Lower Respiratory Tract Panel | OpGen | Respiratory | mecA, TEM, CTX-M, KPC, NDM, OXA-23, OXA-24, OXA-48, OXA-58, VIM | 4.5 h | Yes |
| Rapid Antimicrobial Susceptibility Tests (RASTs): | |||||
| VITEK Reveal | bioMérieux | Positive blood culture | Rapid AST | 5.5 h | Yes |
| Accelerate PhenoTest BC Kit | Accelerate Diagnostics | Positive blood culture | Rapid AST in addition to identification of various bacteria and yeast | 7 h (AST) | Yes |
| ASTar | Q-linea | Positive blood culture | Rapid AST | 6 h | Yes |
| Next Generation Phenotyping (NGP) Panels | Selux Diagnostics | All samples | NGP-GP: Rapid AST for 14 gram-positive antimicrobial agents NGP-GN: Rapid AST for 23 gram-negative antimicrobial agents | 5-6 h | Yes |
| AST, antimicrobial susceptibility testing; BC-GP Panel, blood culture gram-positive panel; BC-GN Panel, blood culture gram-negative panel; BCID-GP Panel, blood culture identification gram-positive panel; BCID-GN Panel, blood culture identification gram-negative panel; CTX-M, Cefotaxime bet-lactamase; EUA, emergency use authorization; IMP, imipenemase-type metallo-beta-lactamase; KPC, Klebsiella pneumoniae carbapenemase; NDM, New Delhi metallo-beta-lactamase; OXA, oxacillinase; NGP-GN, next-generation phenotyping gram-negative panel; NGP-GP, next-generation phenotyping gram-positive panel; MRSA, methicillin-resistant Staphylococcus aureus; RDT, rapid diagnostic test; SHV, sulfhydryl variable beta-lactamases; TAT, turnaround time; VIM, Verona integron–encoded metallo-beta-lactamase. a FDA cleared only. | |||||
Rapid Antimicrobial Susceptibility Testing
The need for novel methods to reduce time to antimicrobial susceptibility results is urgent as the current gold-standard method (ie, broth microdilution [BMD]) can take days to weeks to produce a result. The BMD method determines the MIC of the bacterial strain, which helps guide treatment decisions. Although the BMD method is the gold standard, the Kirby-Bauer disc diffusion method has also been used commonly for AST in clinical microbiological laboratories since its development in 1940.107,108 This method involves culturing bacteria on an agar plate and exposing them to paper discs impregnated with different antibiotics. After incubation, the zones of inhibition are measured to determine whether bacteria are susceptible or resistant to antibiotics tested. The BMD method has an advantage over the Kirby-Bauer disc diffusion method in that the quantitative MIC values allow us to assess the degree of resistance or susceptibility.109-111 However, these traditional methods can take a long time for the results to be available (ie, days to weeks from positive culture).112,113
Fortunately, the availability of RASTs may help to significantly shorten the time to result from days to hours. For example, the FDA-cleared Accelerate PhenoTest BC system provides a shorter turnaround time for identification and AST (to approximately 2 and 7 hours direct from positive blood culture, respectively) by using morphokinetic cellular analysis.114 A multicenter retrospective observational study compared clinical and antimicrobial stewardship metrics before and after implementation of Accelerate PhenoTest BC Kit (AXDX) in patients with BSIs where 854 patients (435 pre-AXDX, 419 post-AXDX) were included. This study found the median time to optimal therapy was 17.2 hours shorter in the post-AXDX arm (23.7 hours) compared with the pre-AXDX arm (40.9 hours; P<0.0001), and LOS was shorter in the post-AXDX arm (5.4 vs 6.4 days; P=0.03) among patients with gram-negative bacteremia.115 Another RAST example is the FDA-cleared Q-linea ASTar, which uses high-speed, high-resolution optical detection and an image analysis algorithm to continuously evaluate the collected images to quantify accumulating bacterial biomass. One study evaluated the time to result and impact on antimicrobial therapy of ASTar compared with VITEK 2 from short-term culture.116 The categorical agreement was found to be similar to standard procedure (95.6%). The AST results were provided significantly faster (~15 hours) with ASTar compared with VITEK 2. However, the percentage of patients who will profit from ASTar in a low-resistance setting is limited, and it is unclear whether a change of therapy 15 hours sooner will have a significant impact on clinical outcomes.116 Another new FDA-cleared method for RAST is called Next Generation Phenotyping (NGP) used by Selux. This novel RAST system uses 2 sequential assays performed on all wells of doubling dilution series to determine MICs. A recent study evaluated the performance of the Selux NGP system compared with BMD. High agreement between both methods was demonstrated (>95%), and the average time to result (from time of assay start to time of MIC result) was 5.65 hours.117 Lastly, the FDA-cleared VITEK Reveal system represents another RAST method that uses volatile metabolites analysis for RAST of gram-negative pathogens from positive blood culture. A study comparing VITEK Reveal and BMD found relatively high categorical and essential agreement (both >90%) except with some specific antibiotic-pathogen combinations.118 Several other methods of RAST are being developed, including the QuantaMatrix dRAST system, which is a novel rapid AST system that uses microfluidic technology and microscopic imaging technology.119 Table 4 includes a summary and comparison of selected systems for RAST.
Conclusion
The rapid advancement of RDTs and RASTs can play a crucial role in identifying causative pathogens, testing for susceptibility, and detecting resistance markers quickly. RDTs can shorten the time between specimen collection and diagnostic results compared with traditional culture methods. In addition, non–culture-based methods eliminate the need for long incubation, which further reduces the time to results. Implementing these tests allows for rapid initiation of appropriate therapy and avoidance of inappropriate antimicrobial use, which contributes toward the development of antimicrobial resistance and adverse events. RDTs and RASTs can also play a key role in antimicrobial de-escalation, which is an important aspect of ASPs. RDTs and RASTs showed promising results in terms of accuracy and turnaround time. Moving forward, efforts toward eliminating implementation barriers should be emphasized to facilitate use in clinical microbiology laboratories.
References
- Eubank TA, Long SW, Perez KK. J Infect Dis. 2020;222(suppl 2):S103-S109.
- WHO Bacterial Priority Pathogens List, 2024. Accessed September 12, 2024. https://www.who.int/publications/i/item/9789240093461
- Curren EJ, Lutgring JD, Kabbani S, et al. Clin Infect Dis. 2022;74(4):723-728.
- CDC. Core elements of hospital antibiotic stewardship programs. Accessed September 12, 2024. https://bit.ly/3Tzjz3P-IDSE
- CDC, Nemhauser JB. CDC Yellow Book 2024: Health Information for International Travel. Accessed September 4, 2024. https://doi.org/10.1093/oso/9780197570944.001.0001
- Bouzid D, Zanella MC, Kerneis S, et al. Clin Microbiol Infect. 2021;27(2):182-191.
- Schmitz JE, Stratton CW, Persing DH, et al. J Clin Microbiol. 2022;60(10):e02446-21.
- Hanson KE, Azar MM, Banerjee R, et al. Clin Infect Dis. 2020;71(10):2744-2751.
- FDA. In vitro diagnostics EUAs—molecular diagnostic tests for SARS-CoV-2. Department of Health and Human Services, FDA; 2024.
- Kalil AC, Metersky ML, Klompas M, et al. Clin Infect Dis. 2016;63(5):e61-e111.
- Metlay JP, Waterer GW, Long AC, et al. Am J Respir Crit Care Med. 2019;200(7):e45-e67.
- Markussen DL, Serigstad S, Ritz C, et al. JAMA Network Open. 2024;7(3):e240830.
- Poole S, Tanner AR, Naidu VV, et al. J Infect. 2022;85(6):625-633.
- Clark TW, Lindsley K, Wigmosta TB, et al. J Infect. 2023;86(5):462-475.
- Rogan M. Respiratory infections, acute. International Encyclopedia of Public Health. Published online October 24, 2016. doi:10.1016/B978-0-12-803678-5.00383-0
- Jin X, Ren J, Li R, et al. EClinicalMedicine. 2021;37:100986.
- Safiri S, Mahmoodpoor A, Kolahi AA, et al. Front Public Health. 2023;10:1028525.
- GBD 2016 Lower Respiratory Infections Collaborators. Lancet Infect Dis. 2018;18(11):1191-1210.
- Almirall J, Bolíbar I, Balanzó X, et al. Eur Respir J. 1999;13(2):349-355.
- Tazinya AA, Halle-Ekane GE, Mbuagbaw LT, et al. BMC Pulm Med. 2018;18(1):7.
- MacVane SH, Oppermann N, Humphries RM. J Clin Microbiol. 2020;58(11):e01468-20.
- Calderaro A, Buttrini M, Farina B, et al. Microorganisms. 2022;10(9):1856.
- Kuti EL, Patel AA, Coleman CI. J Crit Care. 2008;23(1):91-100.
- Poole S, Townsend J, Wertheim H, et al. Eur J Clin Microbiol Infect Dis. 2021;40(2):429-434.
- Sur DKC, Plesa ML. Am Fam Physician. 2022;106(6):628-636.
- Grief SN. Prim Care. 2013;40(3):757-770.
- Saleri N, Ryan ET. Respiratory infections. Travel Medicine. 4th ed. Elsevier; 2019:527-537. doi:10.1016/B978-0-323-54696-6.00059-8
- Mandell GL, Bennett JE, Dolin R, et al. Arch Pathol Lab Med. 1997;121(8):908.
- Havers FP, Hicks LA, Chung JR, et al. JAMA Netw Open. 2018;1(2):e180243.
- Poritz MA, Blaschke AJ, Byington CL, et al. PLoS One. 2011;6(10):e26047.
- Popowitch EB, O’Neill SS, Miller MB. J Clin Microbiol. 2013;51(5):1528-1533.
- Pierce VM, Elkan M, Leet M, et al. J Clin Microbiol. 2012;50(2):364-371.
- VERIGENE Respiratory Pathogens Flex Panel. Package insert. Accessed September 5, 2024. https://bit.ly/3ZoRXSR-IDSE
- Luminex Corporation. Accessed September 5, 2024. https://bit.ly/3XwfzSY-IDSE
- Leber AL, Everhart K, Daly JA, et al. J Clin Microbiol. 2018;56(6):e01945-17.
- Leber AL, Lisby JG, Hansen G, et al. J Clin Microbiol. 2020;58(5):e00155-120.
- Visseaux B, Le Hingrat Q, Collin G, et al. J Clin Microbiol. 2020;58(8):e00630-20.
- van Asten SAV, Boers SA, de Groot JDF, et al. BMC Microbiol. 2021;21:236.
- Jarrett J, Uhteg K, Forman MS, et al. J Clin Virol. 2021;135:104737.
- NxTAG Respiratory Pathogen Panel. Package insert. Accessed September 5, 2024. https://www.biomed-global.com/wp-content/uploads/2021/08/NxTAG-RPP-CAN-IVD-Catalogue-1.pdf
- Chen JHK, Yip CCY, Chan JFW, et al. J Clin Microbiol. 2020;58(8):e00936-20.
- Beckmann C, Hirsch HH. J Med Virol. 2016;88(8):1319-1324.
- Popowitch EB, Kaplan S, Wu Z, et al. Microbiol Spectr. 2022;10(4): e0124822.
- Chan KH, To KKW, Li PTW, et al. Adv Virol. 2017;2017:1324276.
- Nijhuis RHT, Guerendiain D, Claas ECJ, et al. J Clin Microbiol. 2017;55(6):1938-1945.
- Babady NE, England MR, Jurcic Smith KL, et al. J Clin Microbiol. 2018;56(2):e01658-17.
- Kung HC, Hoyert DL, Xu J, et al. Natl Vital Stat Rep. 2008;56(10):1-120.
- Ramirez JA, Wiemken TL, Peyrani P, et al. Clin Infect Dis. 2017;65(11):1806-1812.
- Ruuskanen O, Lahti E, Jennings LC, et al. Lancet. 2011;377(9773):1264-1275.
- Huang DT, Yealy DM, Filbin MR, et al. N Engl J Med. 2018;379(3):236-249.
- Ruuskanen O, Lahti E, Jennings LC, et al. Lancet. 2011;377(9773):1264-1275.
- Hanson KE, Azar MM, Banerjee R, et al. Clin Infect Dis. 2020;71(10):2744-2751.
- Buchan BW, Windham S, Balada-Llasat JM, et al. J Clin Microbiol. 2020;58(7):e00135-20.
- Webber DM, Wallace MA, Burnham CA, et al. J Clin Microbiol. 2020;58(7):e00343-20.
- Ginocchio CC, Garcia-Mondragon C, Mauerhofer B, et al. Eur J Clin Microbiol Infect Dis. 2021;40(8):1609-1622.
- Zacharioudakis IM, Zervou FN, Dubrovskaya Y, et al. Int J Infect Dis. 2021;104:354-360.
- Monard C, Pehlivan J, Auger G, et al. Crit Care. 2020;24:434.
- Collins ME, Popowitch EB, Miller MB. J Clin Microbiol. 2020;58(5):e02013-19.
- Klein M, Bacher J, Barth S, et al. J Clin Microbiol. 2021;59(3):e02497-20.
- Peiffer-Smadja N, Bouadma L, Mathy V, et al. Crit Care. 2020;24(1):366.
- Pickens C, Wunderink RG, Qi C, et al. Diagn Microbiol Infect Dis. 2020;98(4):115179.
- Hale R, Crowley P, Dervisevic S, et al. Viruses. 2021;13(10):2028.
- Tan CCS, Ko KKK, Chen H, et al. Nat Microbiol. 2023;8(5):973-985.
- Viscoli C. Virulence. 2016;7(3):248-251.
- Santella B, Folliero V, Pirofalo GM, et al. Antibiotics (Basel). 2020;9(12):851.
- Timsit JF, Ruppé E, Barbier F, et al. Intensive Care Med. 2020;46(2):266-284.
- Murray PR, Masur H. Crit Care Med. 2012;40(12):3277-3282.
- Brun-Buisson C, Doyon F, Carlet J, et al. JAMA. 1995;274(12):968-974.
- Dunbar SA, Gardner C, Das S. Front Cell Infect Microbiol. 2022;12:859935.
- Peri AM, Chatfield MD, Ling W, et al. Clin Infect Dis. 2024;79(2):502-515.
- Timbrook TT, Morton JB, McConeghy KW, et al. Clin Infect Dis. 2017;64(1):15-23.
- Pliakos EE, Andreatos N, Shehadeh F, et al. Clin Microbiol Rev. 2018;31(3):e00095-e00117.
- Brosh-Nissimov T, Tzur A, Grupel D, et al. Ann Clin Microbiol Antimicrob. 2023;22:62.
- Kumar A. Curr Infect Dis Rep. 2010;12(5):336-344.
- Claeys KC, Schlaffer K, Smith R, et al. Clin Infect Dis. 2021;73(6):1103-1106.
- Gill CM, Kenney RM, Hencken L, et al. Diagn Microbiol Infect Dis. 2019;95(2):162-165.
- Bassetti M, Righi E, Ansaldi F, et al. Intensive Care Med. 2015;41(9):1601-1610.
- Lamoth F, Akan H, Andes D, et al. Clin Infect Dis. 2021;72(suppl 2):S102-S108.
- Walker T, Dumadag S, Lee CJ, et al. J Clin Microbiol. 2016;54(7):1789-1796.
- Ward C, Stocker K, Begum J, et al. Eur J Clin Microbiol Infect Dis. 2015;34(3):487-496.
- Huang TD, Melnik E, Bogaerts P, et al. J Clin Microbiol. 2019;57(2):e01597-e01618.
- McCarty TP, Cumagun P, Meeder J, et al. Microbiol Spectr. 2023;11(1):e04092-22.
- Samuel L. J Clin Microbiol. 2023;61(7):e00231-21.
- Krifors A, Ullberg M, Castegren M, et al. J Fungi (Basel). 2022;8(1):86.
- Nguyen MH, Clancy CJ, Pasculle AW, et al. Ann Intern Med. 2019;170(12):845-852.
- Banerjee R, Komarow L, Virk A, et al. Clin Infect Dis. 2021;73(1):e39-e46.
- Kakoullis L, Papachristodoulou E, Chra P, et al. Antibiotics (Basel). 2021;10(4):415.
- Sawa T, Kooguchi K, Moriyama K. J Intensive Care. 2020;8:13.
- Dadgostar P. Infect Drug Resist. 2019;12:3903-3910.
- Tamma PD, Heil EL, Justo JA, et al. Accessed September 12, 2024. https://www.idsociety.org/practice-guideline/amr-guidance/
- Jenkins S, Ledeboer NA, Westblake LF, et al. J Clin Microbio. 2020;58(7):e00344.
- Dortet L, Tandé D, de Briel D, et al. J Antimicrob Chemother. 2018;73(9):2352-2359.
- Yoo IY, Huh K, Shim HJ, et al. Int J Infect Dis. 2020;95:326-331.
- Richter SS, Marchaim D. Virulence. 2016;8(4):417-426.
- Lindblom A, Karami N, Magnusson T, et al. Eur J Clin Microbiol Infect Dis. 2018;37(8):1491-1497.
- Rottier WC, Bamberg YRP, Dorigo-Zetsma JW, et al. Clin Infect Dis. 2015;60(11):1622-1630
- Parente DM, Cunha CB, Mylonakis E, et al. Clin Infect Dis. 2018;67(1):1-7.
- Willis C, Allen B, Tucker C, et al. Am J Health Syst Pharm. 2017;74(21):1765-1773.
- Smith MN, Erdman MJ, Ferreira JA, et al. J Crit Care. 2017;38:168-171.
- Smith MN, Brotherton AL, Lusardi K, et al. Ann Pharmacother. 2019;53(6):627-638.
- Carr AL, Daley MJ, Givens Merkel K, et al. Pharmacotherapy. 2018;38(12):1216-1228.
- Butler-Laporte G, De L’étoile-Morel S, Cheng MP, et al. J Infect. 2018;77(6):489-495.
- Mergenhagen KA, Starr KE, Wattengel BA, et al. Clin Infect Dis. 2020;71(5):1142-1148.
- Shaw R, Zander A, Ronnie T, et al. Open Forum Infect Dis. 2024;11(7):ofae408.
- Page A, O’Rourke S. Brennan M, et al. I J Med Sci. 2017;186(4):995-996.
- Guarino M, Perna B, Cesaro AE. J Clin Med. 2023;12(9):3188.
- Clinical and Laboratory Standards Institute (CLSI). M02. Performance Standards for Antimicrobial Disk Susceptibility Tests. 13th ed. Clinical and Laboratory Standards Institute; 2018.
- The European Committee on Antimicrobial Susceptibility Testing (EUCAST): clinical breakpoints and dosing of antibiotics. Breakpoint tables for interprettion of MICS and Zone diaameters. Accessed September 6, 2024. https://www.eucast.org/clinical_breakpoints
- Clinical and Laboratory Standards Institute (CLSI). M07. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically. 11th ed. Clinical and Laboratory Standards Institute; 2018.
- Golus J, Sawicki R, Widelski J, et al. J Appl Microbiol. 2016;121(5):1291-1299.
- Kowalska-Krochmal B, Dudek-Wicher R. Pathogens. 2021;10(2):165.
- Smith KP, Kirby JE. Rapid susceptibility testing methods. Clin Lab Med. 2019;39(3):333-344.
- Khan ZA, Siddiqui MF, Park S. Diagnostics (Basel). 2019;9(2):49.
- Brosh-Nissimov T, Tzur A, Grupel D, et al. Ann Clin Microbiol Antimicrob. 2023;22:62.
- Esse J, Träger J, Valenza G, et al. J Clin Microbiol. 2023;61(11):e0054923.
- Baker KR, Flentie K, Spears BR, et al. J Clin Microbiol. 2024;62(1):e00546-23.
- Antonelli A, Cuffari S, Casciato B, et al. Diagn Microbiol Infect Dis. 2024;110(4):116503.
- Rosselin M, Prod’hom G, Greub G, et al. Microorganisms. 2022;10(6):1212.
- Wong AYW, Johnsson ATA, özenci V. Microbiol Spectrum. 2022;10(2):e02107-21.
Copyright © 2024 McMahon Publishing, 545 West 45th Street, New York, NY 10036. Printed in the USA. All rights reserved, including the right of reproduction, in whole or in part, in any form.

Download to read this article in PDF document:![]() RDTs and RASTs: The Current Landscape
RDTs and RASTs: The Current Landscape