In a recent article appearing in this publication, I provided an overview on the growing importance of rapid diagnostics for infectious disease care. That review focused broadly on the myriad clinical and laboratory scenarios through which the ID field is being transformed by diagnostics with improved turnaround times. These scenarios range from point-of-care (POC) testing within clinics to the in-lab identification of microbial isolates and their antibiotic susceptibility profiles.1 Because this issue of Infectious Disease Special Edition focuses precisely on the theme of rapid ID testing, this follow-up article takes a deeper dive into one of its most impactful manifestations: rapid diagnostic tests (RDTs) for community-acquired infections, particularly assays that entail nucleic acid amplification tests (NAATs). Various molecular RDTs have entered clinical care over the past decade, with their targeted detection of pathogen-specific DNA and/or RNA (the latter requiring an initial reverse transcription step). Respiratory tract infections (RTIs), especially upper RTIs (URTIs), represent the most common use case for molecular RDTs, even more so since the emergence of SARS-CoV-2. These tests likewise stand as an important progression from the immunodiagnostic RDTs that preceded them. The present discussion provides a combined perspective on the regulatory, technical, and clinical factors driving the expansion of molecular RDTs for respiratory infections, including examples of on-market products in the United States.
What’s in a Name?
The term RDT can be, and has been, applied loosely across different clinical and laboratory contexts. In 2009, for instance, the European Union Commission on diagnostic devices sought a more rigorous definition for RDTs as “qualitative or semi-quantitative in vitro diagnostic medical devices, used singly or in a small series, which involve non-automated procedures and have been designed to give a fast result.”2 From a 2024 perspective, this definition might seem dated, especially with the advent of the molecular technologies that are the focus of the present review. Likewise, the terms RDT and POC testing are often applied interchangeably, as many RDTs are designed specifically with POC use in mind. That said, other assays frequently classified under the RDT umbrella can be performed within dedicated laboratory facilities (typically ones where specimens can arrive shortly after collection). For this article, RDTs will be treated as diagnostics with the following characteristics, regardless of whether they are performed at the POC or in a laboratory:
- short performance times (=1-2 hours, but often far less);
- ease of use, with minimal manual steps and potential accessibility to non-laboratorians;
- random-access capability, negating the need for high-order batching and fixed testing schedules; and
- real-time applicability to clinical decision making.3
RDTs are developed and marketed commercially, and the settings in which they may be deployed are affected by pertinent regulations. Despite any looseness around the term itself, key definitions that govern RDT usage carry formal, legal weight in the United States, namely, the complexity levels stipulated by the Clinical Laboratory Improvement Amendments (CLIA). When the FDA grants in vitro diagnostic status to a new assay, the test is typically assigned a CLIA complexity level of high, moderate, or waived (based on 7 scored criteria covering training, operation, and interpretation).4 High- and moderate-complexity tests must be performed within CLIA-certified laboratory environments, under stringent requirements for quality assurance, licensing, and inspection. By contrast, waived complexity tests cover the simplest analytical procedures with the lowest possibility of erroneous results. Only waived assays, following appropriate clinical trials, may be marketed for direct use by providers at the POC or by patients themselves at home or in other point-of-need scenarios. In total, therefore, RDTs encompass a combination of CLIA-waived diagnostics, together with some non-waived assays performed in the lab. Diagnostic regulations vary with location, and RDTs with non-waived status in the United States are often employed more flexibility in global settings with limited laboratory infrastructure. For other RDTs, in turn, the sole focus may be international markets.
From Terminology to Technology
Although this CLIA complexity framework has remained in place for nearly 30 years, RDT technologies themselves have evolved considerably. Until recently, microbial RDTs were based exclusively on immunodiagnostic methods, most commonly the lateral flow assay (LFA). This technique relies on the capture of pathogen-specific analytes by antibodies immobilized on a membrane, through which a liquid/suspended diagnostic sample flows via capillary action.5 These analytes typically represent pathogen-specific proteins or carbohydrates (for rapid antigen tests), although they may also represent patient-derived antibodies against a pathogen (for rapid serological tests). A second antibody-binding event, in turn, generates the LFA readout through a conjugated chromophore or fluorophore. Over the years, LFAs have been adapted into RDTs for diverse specimen types and pathogens, including viral,6-8 bacterial,9-11 fungal,12,13 and parasitic agents.14,15 Many such assays still represent critical components of diagnostic care, across diverse infections and global settings. LFAs are inherently rapid, and they are readily adapted into straightforward protocols for non-laboratorians.
With their elegance in simplicity, a variety of LFAs have been granted waived complexity status for both microbial detection and non-ID diagnostic purposes.16 By contrast, for molecular RDTs introduced more recently, technical simplicity may not be the first description these platforms evoke. The technology behind the assays is anything but simple; in fact, they represent impressive feats of biomedical engineering. At the same time, a defining feature of molecular RDTs is that their processes are carried out within small, self-enclosed instruments. To the perspective of the end user, the protocol simply involved inoculation of a diagnostic specimen into a test cartridge and insertion of that cartridge into a benchtop device. The automated processes within these devices make rapid NAATs possible, facilitating sample-to-result analyses with greater sensitivity—at least for many pathogens—than can be achieved through immunoassays. Critically, the steps of a molecular RDT entail more than just targeted amplification of nucleic acid. Whether traditional or rapid, NAATs typically entail 3 sequential components that together dictate assay performance—namely, nucleic acid extraction, target amplification, and amplicon detection.17 These steps must be integrated seamlessly for an RDT, a need that is addressed by one of the defining features of such assays, automated microfluidics. Fundamentally, this internal liquid handling allows complex laboratory technique to be translated outside dedicated lab settings.
As to the technical details of extraction, amplification, and detection, a variety of strategies are employed across RDT platforms. For instance, although the terms nucleic acid amplification and polymerase chain reaction (PCR) are often treated synonymously, different RDTs employ various amplification chemistries. Several common platforms use PCR, including the bioMérieux/Biofire FilmArray/SpotFire,18 Roche/GenMark cobas ePlex,19 Diasorin/Luminex LIAISON PLEX/VERIGENE,20 Roche cobas liat,21 Qiagen QIAstat,22 and Cepheid Gene- Xpert.23 The small liquid volumes manipulated by such instruments are also important for optimizing turnaround, as they minimize the time required for thermocycling (with total amplification times as low as about 30 minutes). However, other RDT platforms exploit non-PCR isothermal amplification techniques, including nicking endonuclease amplification24 (Abbott ID Now), the helicase-dependent amplification25 (QuidelOrtho Solana), and loop-mediated isothermal amplification26 (Meridian Alethia).
Methods for amplicon detection are equally diverse, highly influenced by the upstream amplification step. While the numerous molecular details are beyond the scope of this review, different RDTs generate readable signals through strategies that include fluorescence18,21-23 (both probe-based and non–probe-based), other optical properties,27 electrochemical detection,19 or even turbidity.26 Whereas some instruments employ a single readout at the conclusion of amplification, others monitor the amplification signal throughout the course of the reaction. The latter include assays based on the real-time PCR (qPCR) reaction, with probe-based fluorescent readouts.28 In theory, the cycle threshold (Ct) values generated by qPCR can facilitate absolute pathogen quantification—with inclusion of appropriate calibrators—although rapid qPCR platforms only use the technique to generate categorical detected/not detected results. (For some qPCR-based RDT assays, in fact, the measured Ct values are blocked from the user).
The fluorescence-based approach of qPCR also highlights another common feature of molecular RDTs: the ability to evaluate for multiple targets simultaneously. By varying the color of fluorophore associated with an amplicon-specific probe, a rapid qPCR assay can differentiate between several amplicons (typically up to 4) as part of a multiplex amplification protocol.28 These targets typically represent different pathogens that elicit overlapping infectious pictures. The desire for diagnostic breadth can extend to even more targets, serving as the motivation for highly multiplex “syndromic” RDTs29 (eg, the FilmArray/SpotFire, ePlex, and LIAISON PLEX/VERIGENE, and QIAstat). And these infectious syndromes now covered by these platforms include URTIs,30 lower RTIs/pneumonia,31 gastroenteritis/colitis,32 central nervous system infections,33 bloodstream infections,34,35 and bone/joint infections.36 To evaluate for more than a dozen targets, these platforms must overcome the limitations of spectral overlap, as only so many fluorophores can be distinguished from one another simultaneously. In broad strokes, they achieve higher-order multiplexing by detecting amplicons across spatially discriminant locations within the testing device. Again, the fact that these hands-off capabilities are possible is a testament to the level of engineering that underpins molecular RDTs.
Molecular RDTs in Clinical Context
The technical features of these instruments allow them to meet the first 3 criteria for a successful RDT noted above (short turnaround time, ease of use, and random access). But as to the fourth criteria—the ability to influence clinical decisions in real time—the value of a rapid assay is also determined by epidemiological variables and standards of available care. RTIs illustrate this dynamic, as both the most common infectious disease and the most widespread application of RDTs. When evaluating a patient with a suspected RTI, one of the first questions any clinician must assess is whether the upper and/or lower respiratory tract is involved. If lower respiratory involvement is suspected, not only does it raise the possible need for elevated care but expands the differential diagnosis (eg, to include agents of bacterial pneumonia). However, even for purely upper respiratory involvement, the potential etiologic agents are diverse, and they can demonstrate substantial symptomatic overlap with one another.37 Viral pathogens are overwhelmingly the most common, and they include influenza A/B, SARS-CoV-2, other coronaviruses, enteroviruses (including rhinoviruses), respiratory syncytial virus (RSV), parainfluenza, metapneumovirus, adenovirus, and bocavirus. For many patients evaluated for URTIs (especially children), streptococcal pharyngitis is also an important consideration.
Given this differential diagnosis (minus SARS-CoV-2), it is instructive to consider a respiratory evaluation from 15+ years ago, when only immunodiagnostic RDTs were available to providers. These included (and still do include) LFAs for influenza A/B and group A Streptococcus, for which routine antiviral and antibiotic agents represent standards of care. The ability of an RDT to guide decision making may be viewed as the individual actionable value of positive and negative results.1 In other words, how effective is the RDT as both a rule-in and rule-out? In the case of respiratory viruses and streptococcal pharyngitis, the analytical limit of detection of immunoassays is inferior to that of viral/bacterial culture and NAATs.a Depending on a patient’s burden of pathogen (which can vary from individual to individual) and when a specimen is collected during an infection, some cases of influenza (now also SARS-CoV-2) and streptococcal pharyngitis are missed by rapid LFAs. With this reduced diagnosed sensitivity—often in the 70% to 90% range, relative to culture/NAATs—these tests can effectively diagnose such infections, but they cannot rigorously exclude them.38-40 In turn, while positive LFA results may prompt a clinician to initiate appropriate therapy or lead to isolation precautions, negative results still require traditional follow-up testing to provide an absolute rule-out. In the meantime, a clinician’s decision whether to treat must remain empiric. If this decision does not match subsequent in-lab data, the theoretical downside is either a treatment delay or (alternatively) a period of unnecessary therapy.
Of note, the preceding discussion is not intended as a negative commentary on the general role of immunodiagnostic RDTs. The ability of these assays to influence medical choices depends on numerous clinical variables. Rapid LFAs continue to make indispensable contributions to routine and specialized care, globally and with various types of infections. Even for URTIs, their rule-in value alone is sufficient to motivate and justify their continued use under many circumstances, such as their recent expansion into at-home testing for SARS-CoV-2.41 In the context of respiratory infections, both the benefits and limitations of immunodiagnostic RDTs help illustrate the expanded value of their molecular successors. Due to the enhanced sensitivity of NAATs to detect specimens of low viral load, molecular RDTs possess superior negative predictive value for excluding URTI pathogens.39,40 This added sensitivity carries certain caveats, including an appreciation that nucleic acid contamination of a testing environment could theoretically lead to false-positive results, particularly relevant for POC testing outside of dedicated labs.42 When carefully performed, however, a notable benefit of these assays is that they may guide clinical decision making broadly, for either positive or negative results.
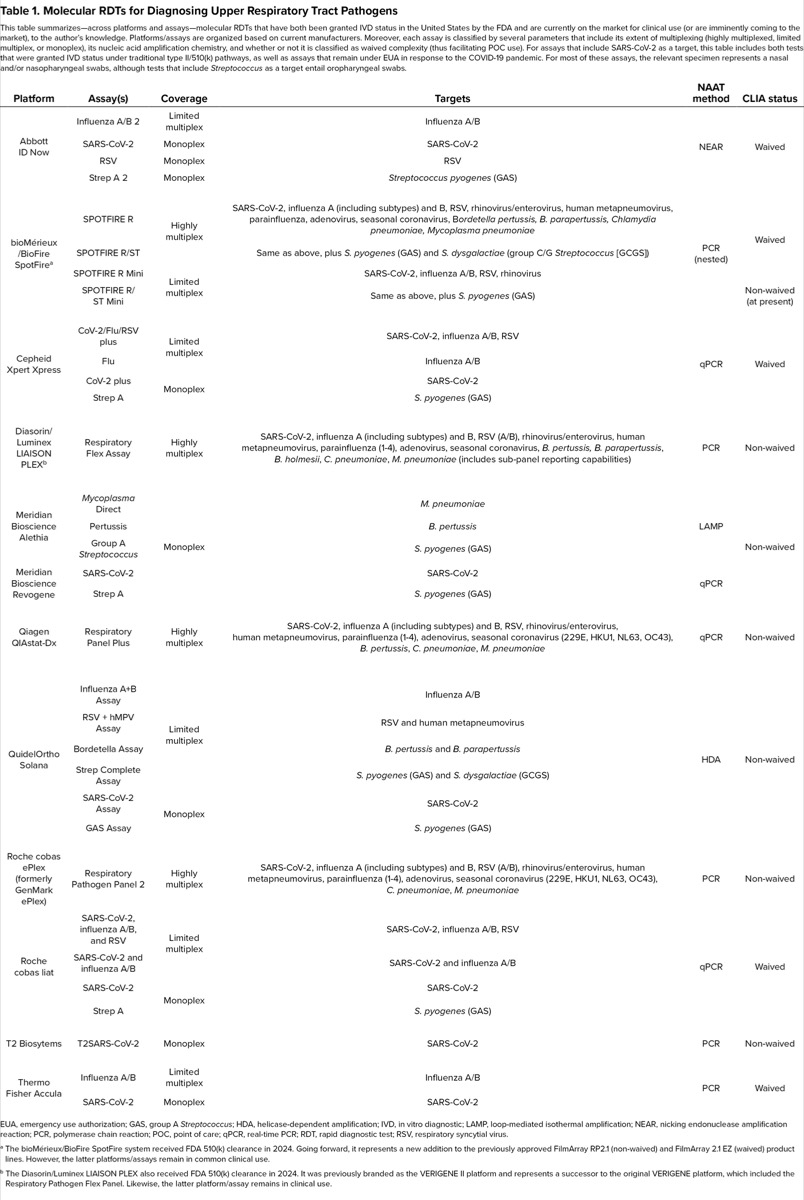
This added value is even more noteworthy considering SARS-CoV-2. As with influenza A/B, this now-common pathogen may be treated with dedicated antiviral therapy to help prevent severe disease. In fact, the availability of targeted management strategies for influenza and SARS-CoV-2 distinguish these viruses among URTI agents. This dynamic is reflected by the development of multitarget assays, including molecular RDTs, that evaluate for these targets in combination. This level of multiplexing is well also within the fluorescent readout capabilities of traditional qPCR, without the need for higher-order multiplexing.28 The Table summarizes molecular RDTs for upper respiratory tract pathogens, currently available in the United States, which are cleared by the FDA. These assays (all commercially developed) are summarized according to several parameters, including their manufacturers, level of multiplexing, specific targets, amplification class, and CLIA status. Of note, the latter implies whether a given assay may be used in POC settings (under a CLIA certificate of waiver) or instead must be conducted within a dedicated laboratory environment. Along with multiplex molecular RDTs, multitarget immunoassay RDTs for upper respiratory pathogens (ie, SARS-COV-2, influenza A/B, and/or RSV) have also entered the US clinical market. Examples include the BD Veritor and Hardy Diagnostics Status assays.
| Table 1. Molecular RDTs for Diagnosing Upper Respiratory Tract Pathogens This table summarizes—across platforms and assays—molecular RDTs that have both been granted IVD status in the United States by the FDA and are currently on the market for clinical use (or are imminently coming to the market), to the author’s knowledge. Platforms/assays are organized based on current manufacturers. Moreover, each assay is classified by several parameters that include its extent of multiplexing (highly multiplexed, limited multiplex, or monoplex), its nucleic acid amplification chemistry, and whether or not it is classified as waived complexity (thus facilitating POC use). For assays that include SARS-CoV-2 as a target, this table includes both tests that were granted IVD status under traditional type II/510(k) pathways, as well as assays that remain under EUA in response to the COVID-19 pandemic. For most of these assays, the relevant specimen represents a nasal and/or nasopharyngeal swabs, although tests that include Streptococcus as a target entail oropharyngeal swabs. | |||||
| Platform | Assay(s) | Coverage | Targets | NAAT method | CLIA status |
|---|---|---|---|---|---|
| Abbott ID Now | Influenza A/B 2 | Limited multiplex | Influenza A/B | NEAR | Waived |
| SARS-CoV-2 | Monoplex | SARS-CoV-2 | |||
| RSV | Monoplex | RSV | |||
| Strep A 2 | Monoplex | Streptococcus pyogenes (GAS) | |||
| bioMérieux /BioFire SpotFirea | SPOTFIRE R | Highly multiplex | SARS-CoV-2, influenza A (including subtypes) and B, RSV, rhinovirus/enterovirus, human metapneumovirus, parainfluenza, adenovirus, seasonal coronavirus, Bordetella pertussis, B. parapertussis, Chlamydia pneumoniae, Mycoplasma pneumoniae | PCR (nested) | Waived |
| SPOTFIRE R/ST | Same as above, plus S. pyogenes (GAS) and S. dysgalactiae (group C/G Streptococcus [GCGS]) | ||||
| SPOTFIRE R Mini | Limited multiplex | SARS-CoV-2, influenza A/B, RSV, rhinovirus | |||
| SPOTFIRE R/ST Mini | Same as above, plus S. pyogenes (GAS) | Non-waived (at present) | |||
| Cepheid Xpert Xpress | CoV-2/Flu/RSV plus | Limited multiplex | SARS-CoV-2, influenza A/B, RSV | qPCR | Waived |
| Flu | Influenza A/B | ||||
| CoV-2 plus | Monoplex | SARS-CoV-2 | |||
| Strep A | S. pyogenes (GAS) | ||||
| Diasorin/Luminex LIAISON PLEXb | Respiratory Flex Assay | Highly multiplex | SARS-CoV-2, influenza A (including subtypes) and B, RSV (A/B), rhinovirus/enterovirus, human metapneumovirus, parainfluenza (1-4), adenovirus, seasonal coronavirus, B. pertussis, B. parapertussis, B. holmesii, C. pneumoniae, M. pneumoniae (includes sub-panel reporting capabilities) | PCR | Non-waived |
| Meridian Bioscience Alethia | Mycoplasma Direct | Monoplex | M. pneumoniae | LAMP | Non-waived |
| Pertussis | B. pertussis | ||||
| Group A Streptococcus | S. pyogenes (GAS) | ||||
| Meridian Bioscience Revogene | SARS-CoV-2 | SARS-CoV-2 | qPCR | ||
| Strep A | S. pyogenes (GAS) | ||||
| Qiagen QIAstat-Dx | Respiratory Panel Plus | Highly multiplex | SARS-CoV-2, influenza A (including subtypes) and B, RSV, rhinovirus/enterovirus, human metapneumovirus, parainfluenza (1-4), adenovirus, seasonal coronavirus (229E, HKU1, NL63, OC43), B. pertussis, C. pneumoniae, M. pneumoniae | qPCR | Non-waived |
| QuidelOrtho Solana | Influenza A+B Assay | Limited multiplex | Influenza A/B | HDA | Non-waived |
| RSV + hMPV Assay | RSV and human metapneumovirus | ||||
| Bordetella Assay | B. pertussis and B. parapertussis | ||||
| Strep Complete Assay | S. pyogenes (GAS) and S. dysgalactiae (GCGS) | ||||
| SARS-CoV-2 Assay | Monoplex | SARS-CoV-2 | |||
| GAS Assay | S. pyogenes (GAS) | ||||
| Roche cobas ePlex (formerly GenMark ePlex) | Respiratory Pathogen Panel 2 | Highly multiplex | SARS-CoV-2, influenza A (including subtypes) and B, RSV (A/B), rhinovirus/enterovirus, human metapneumovirus, parainfluenza (1-4), adenovirus, seasonal coronavirus (229E, HKU1, NL63, OC43), C. pneumoniae, M. pneumoniae | PCR | Non-waived |
| Roche cobas liat | SARS-CoV-2, influenza A/B, and RSV | Limited multiplex | SARS-CoV-2, influenza A/B, RSV | qPCR | Waived |
| SARS-CoV-2 and influenza A/B | SARS-CoV-2 and influenza A/B | ||||
| SARS-CoV-2 | Monoplex | SARS-CoV-2 | |||
| Strep A | S. pyogenes (GAS) | ||||
| T2 Biosytems | T2SARS-CoV-2 | Monoplex | SARS-CoV-2 | PCR | Non-waived |
| Thermo Fisher Accula | Influenza A/B | Limited multiplex | Influenza A/B | PCR | Waived |
| SARS-CoV-2 | Monoplex | SARS-CoV-2 | |||
| EUA, emergency use authorization; GAS, group A Streptococcus; HDA, helicase-dependent amplification; IVD, in vitro diagnostic; LAMP, loop-mediated isothermal amplification; NEAR, nicking endonuclease amplification reaction; PCR, polymerase chain reaction; POC, point of care; qPCR, real-time PCR; RDT, rapid diagnostic test; RSV, respiratory syncytial virus. a The bioMérieux/BioFire SpotFire system received FDA 510(k) clearance in 2024. Going forward, it represents a new addition to the previously approved FilmArray RP2.1 (non-waived) and FilmArray 2.1 EZ (waived) product lines. However, the latter platforms/assays remain in common clinical use. b The Diasorin/Luminex LIAISON PLEX also received FDA 510(k) clearance in 2024. It was previously branded as the VERIGENE II platform and represents a successor to the original VERIGENE platform, which included the Respiratory Pathogen Flex Panel. Likewise, the latter platform/assay remains in clinical use. | |||||
With the availability of such testing, however, a follow-up question naturally arises: What other common respiratory pathogens should be covered under the routine capabilities of molecular RDTs? Unfortunately, the considerations here become far more variable, and one must consider the corresponding issue of “actionability.” More diagnostic data do not necessarily represent better data if this information cannot guide patient management and outcomes, a common refrain under the objective of diagnostic stewardship. Therefore, for what other pathogens does an explicit identification (or rule-out) carry actionable value? For an otherwise healthy adult with low concern for deterioration, the other aforementioned URTI viruses (while microbiologically diverse) typically only entail the same supportive care. In such instances, more extensive multiplex testing, even if obtained rapidly, may not practically influence how that patient is managed. This calculus may swing dramatically under different clinical scenarios and groups of patients. In children and some adults, for instance, RSV carries elevated risks for severe disease and may demand explicit testing, for which, especially in older children, NAATs offer improved sensitivity over an immunodiagnostic assay.43 This dynamic is reflected by the inclusion of RSV in several molecular RDT assays currently on the market (Table). Across different ages, atypical bacterial infections may become more prominent on a differential diagnosis in patients with appropriate clinical pictures (ie, Bordetella, Chlamydia pneumoniae, Legionella, Mycoplasma pneumoniae); these pathogens are likewise distinct also in that they are managed with antibiotics.44 Going further, and taking this reasoning to its logical conclusion, in a patient with sufficient risk factors—or one whose respiratory status is deteriorating and requires admission—an explicit microbiological diagnosis may be indicated regardless of the offending agent. Given these variables, it is perhaps not surprising that outcomes studies evaluating the impact of molecular RDTs have often yielded heterogeneous results, albeit often with general conclusions that inpatient care warrants more aggressive testing.45
In essence, the result becomes a tiered approach to rapid molecular diagnosis. Current methodologies are already sufficient that, when indicated, a truly expansive net can be cast for a patient who presents with a suspected respiratory infection. Instead, the provider must consider what the patient requires in light of their clinical picture.
In many respects, technology itself is not the limiting factor, but discerning and justifying its appropriate use is. Not surprisingly, these same factors also dictate how molecular respiratory testing is billed and reimbursed. Although this point takes us to the limit of the article’s intended scope, it is worth noting that Current Procedural Terminology (CPT) codes for diagnostics in the United States stratify these assays according to the number of multiplexed targets.46 The extent of multiplexing that a payor will reimburse—especially beyond influenza and SARS-CoV-2—is very much affected by a patient’s individual circumstances and setting of care. In fact, the LIAISON PLEX/VERIGENE assay line allows for the flexible release of subsets of targets from a highly multiplexed respiratory panel, to customize the reported data (and billing) around individual patients.
In conclusion, molecular RDTs epitomize the interrelated opportunities and complexities of modern infectious disease care, particularly for managing RTIs. These platforms represent a confluence of clinical need and powerful technology, brought to the forefront by specialized manufacturers and highlighted by an often unpredictable epidemiological landscape. Moreover, the diversity of options available to clinicians and clinical laboratories only promises to grow in the near future, including ambitious goals for at-home molecular testing.47 The field is still working through complex questions about maximizing the added value of molecular RDTs, which remains critically tied to the insights of expert providers. Perhaps the only thing that can be said for certain is that interesting times continue ahead!
aAs a brief aside, one should note that the relative performance of any methodology depends on the pathogen and its course of infection. Infections exist—for instance, certain fungal infections—for which the diagnostic sensitivity of LFAs makes these techniques the gold standard.
References
- Schmitz JE. The need for speed: how emerging rapid diagnostics are transforming infectious disease care. Infectious Disease Special Edition. 2024;28:41-46.
- Commission decision of 27 November 2009 amending decision 2002/364/EC on common technical specifications for in vitro diagnostic medical devices. Official Journal of the European Union. 2009/886/EC. Document 32009D0886. Accessed August 11, 2024. https://bit.ly/ 4cCmsIK-IDSE
- Rabold E, Waggoner J. Rapid Diagnostic tests for infectious diseases. CDC Yellow Book 2024. Accessed August 11, 2024. https://bit.ly/ 3zPB0WE-idse
- FDA. CLIA categorizations. Accessed August 11, 2024. https://www.fda.gov/ medical-devices/ ivd-regulatory-assistance/ clia-categorizations
- Nan X, Yao X, Yang L, et al. Lateral flow assay of pathogenic viruses and bacteria in healthcare. Analyst. 2023;148(19):4573-4590.
- CDC. Rapid influenza diagnostic tests. Accessed August 11, 2024. https://www.cdc.gov/ flu/ professionals/ diagnosis/ clinician_guidance_ridt.htm
- Dinnes J, Sharma P, Berhane S, et al. Rapid, point-of-care antigen tests for diagnosis of SARS-CoV-2 infection. Cochrane Database Syst Rev. 2022;7(7):CD013705.
- Qiu X, Sokoll L, Yip P, et al. Comparative evaluation of three FDA-approved HIV Ag/Ab combination tests using a genetically diverse HIV panel and diagnostic specimens. J Clin Virol. 2017;92:62-68.
- Cohen JF, Bertille N, Cohen R, et al. Rapid antigen detection test for group A Streptococcus in children with pharyngitis. Cochrane Database Syst Rev. 2016;7(7):CD010502.
- Bristow CC, Klausner JD, Tran A. Clinical test performance of a rapid point-of-care syphilis treponemal antibody test: a systematic review and meta-analysis. Clin Infect Dis. 2020;71(suppl 1):S52-S57.
- Kazandjian D, Chiew R, Gilbert GL. Rapid diagnosis of Legionella pneumophila serogroup 1 infection with the Binax enzyme immunoassay urinary antigen test. J Clin Microbiol. 1997;35(4):954-956.
- Abdallah W, Myint T, LaRue R, et al. Diagnosis of histoplasmosis using the MVista Histoplasma galactomannan antigen qualitative lateral flow-based immunoassay: a multicenter study. Open Forum Infect Dis. 2021;8(9):ofab454.
- Tadeo KK, Nimwesiga A, Kwizera R, et al. Evaluation of the diagnostic performance of a semiquantitative cryptococcal antigen point-of-care assay among HIV-infected persons with cryptococcal meningitis. J Clin Microbiol. 2021;59(8):e0086021.
- Kavanaugh MJ, Azzam SE, Rockabrand DM. Malaria rapid diagnostic tests: literary review and recommendation for a quality assurance, quality control algorithm. Diagnostics (Basel). 2021;11(5):768.
- Yanagawa Y, Shimogawara R, Endo T, et al. Utility of the rapid antigen detection test E. histolytica Quik Chek for the diagnosis of Entamoeba histolytica infection in nonendemic situations. J Clin Microbiol. 2020;58(11):e01991-20.
- CDC. Tests granted waived status under CLIA. Accessed July 11, 2024. https://www.cdc.gov/ clia/ docs/ tests-granted-waived-status-under-clia.pdf
- Schmitz JE, Stratton CW, Persing DH, et al. Forty years of molecular diagnostics for infectious diseases. J Clin Microbiol. 2022;60(10):e0244621.
- Poritz MA, Blaschke AJ, Byington CL, et al. FilmArray, an automated nested multiplex PCR system for multi-pathogen detection: development and application to respiratory tract infection. PLoS One. 2011;6(10):e26047.
- Schmitz JE, Tang YW. The GenMark ePlex: another weapon in the syndromic arsenal for infection diagnosis. Future Microbiol. 2018;13(16):1697-1708.
- Huang HS, Tsai CL, Chang J, et al. Multiplex PCR system for the rapid diagnosis of respiratory virus infection: systematic review and meta-analysis. Clin Microbiol Infect. 2018;24(10):1055-1063.
- Chang E, Jeon K, Lee N, et al. Clinical performance of the Roche cobas liat SARS-CoV-2 & influenza A/B assay: a systematic review and meta-analysis. J Clin Virol. 2024;174:105706.
- Parcina M, Schneider UV, Visseaux B, et al. Multicenter evaluation of the QIAstat Respiratory Panel—a new rapid highly multiplexed PCR based assay for diagnosis of acute respiratory tract infections. PLoS One. 2020;15(3):e0230183.
- Cohen DM, Kline J, May LS, et al. Accurate PCR detection of influenza A/B and respiratory syncytial viruses by use of Cepheid Xpert Flu+RSV Xpress Assay in point-of-care settings: comparison to Prodesse ProFlu. J Clin Microbiol. 2018;56(2):e01237-17.
- Srivastava S, Singh P, Malhotra R, et al. Comparison of Abbott ID NOW, a novel isothermal amplification based COVID-19 diagnostic method with RTPCR. J Virol Methods. 2022;304:114521.
- Arbefeville S, Nelson K, Thonen-Kerr E, et al. Prospective postimplementation study of Solana group A streptococcal nucleic acid amplification test vs conventional throat culture. Am J Clin Pathol. 2018;150(4):333-337.
- Ratliff AE, Duffy LB, Waites KB. Comparison of the illumigene Mycoplasma DNA amplification assay and culture for detection of Mycoplasma pneumoniae. J Clin Microbiol. 2014;52(4):1060-1063.
- Cordeiro M, Ferreira Carlos F, Pedrosa P, et al. Gold nanoparticles for diagnostics: advances towards points of care. Diagnostics (Basel). 2016;6(4):43.
- Kralik P, Ricchi M. A basic guide to real time PCR in microbial diagnostics: definitions, parameters, and everything. Front Microbiol. 2017;8:108.
- Ramanan P, Bryson AL, Binnicker MJ, et al. Syndromic panel-based testing in clinical microbiology. Clin Microbiol Rev. 2017;31(1):e00024-17.
- Mills DC, Huder JB, Bloemberg GV, et al. Comparison of three cartridge-based platforms for syndromic testing for respiratory viruses. Diagn Microbiol Infect Dis. 2024;109(3):116308.
- Webber DM, Wallace MA, Burnham CA, et al. Evaluation of the BioFire FilmArray Pneumonia Panel for detection of viral and bacterial pathogens in lower respiratory tract specimens in the setting of a tertiary care academic medical center. J Clin Microbiol. 2020;58(7):e00343-20.
- Khare R, Espy MJ, Cebelinski E, et al. Comparative evaluation of two commercial multiplex panels for detection of gastrointestinal pathogens by use of clinical stool specimens. J Clin Microbiol. 2014;52(10):3667-3673.
- Leber AL, Everhart K, Balada-Llasat JM, et al. Multicenter evaluation of BioFire FilmArray Meningitis/Encephalitis Panel for detection of bacteria, viruses, and yeast in cerebrospinal fluid specimens. J Clin Microbiol. 2016;54(9):2251-61.
- Bork JT, Leekha S, Heil EL, et al. Rapid testing using the Verigene gram-negative blood culture nucleic acid test in combination with antimicrobial stewardship intervention against gram-negative bacteremia. Antimicrob Agents Chemother. 2015;59(3):1588-1595.
- Wojewoda CM, Sercia L, Navas M, et al. Evaluation of the Verigene gram-positive blood culture nucleic acid test for rapid detection of bacteria and resistance determinants. J Clin Microbiol. 2013;51(7):2072-2076.
- Esteban J, Salar-Vidal L, Schmitt BH, et al. Multicenter evaluation of the BIOFIRE Joint Infection Panel for the detection of bacteria, yeast, and AMR genes in synovial fluid samples. J Clin Microbiol. 2023;61(11):e0035723.
- Calderaro A, Buttrini M, Farina B, et al. Respiratory tract infections and laboratory diagnostic methods: a review with a focus on syndromic panel-based assays. Microorganisms. 2022;10(9):1856.
- Shulman ST, Bisno AL, Clegg HW, et al. Clinical practice guideline for the diagnosis and management of group A streptococcal pharyngitis: 2012 update by the Infectious Diseases Society of America. Clin Infect Dis. 2012;55(10):1279-1282.
- Uyeki TM, Bernstein HH, Bradley JS, et al. Clinical practice guidelines by the Infectious Diseases Society of America: 2018 update on diagnosis, treatment, chemoprophylaxis, and institutional outbreak management of seasonal influenza. Clin Infect Dis. 2019;68(6):e1-e47.
- CDC. Overview of testing for SARS-COV-2, the virus that causes COVID-19. Updated March 18, 2024. Accessed August 11, 2024. https://www.cdc.gov/ coronavirus/ 2019-ncov/ hcp/ testing-overview.html
- Katzenschlager S, Brümmer LE, Schmitz S, et al. Comparing SARS-CoV-2 antigen-detection rapid diagnostic tests for COVID-19 self-testing/self-sampling with molecular and professional-use tests: a systematic review and meta-analysis. Sci Rep. 2023;13(1):21913.
- Donato LJ, Myhre NK, Murray MA, et al. Assessment of test performance and potential for environmental contamination associated with a point-of-care molecular assay for group A Streptococcus in an end user setting. J Clin Microbiol. 2019;57(2):e01629-18.
- Onwuchekwa C, Atwell J, Moreo LM, et al. Pediatric respiratory syncytial virus diagnostic testing performance: a systematic review and meta-analysis. J Infect Dis. 2023;228(11):1516-1527.
- Suzuki S, Ishimaru N, Akashi Y, et al. Physicians’ prediction for the assessment of atypical pathogens in respiratory tract infections. J Gen Fam Med. 2020;21(6):226-234.
- Vos LM, Bruning AHL, Reitsma JB, et al. Rapid molecular tests for influenza, respiratory syncytial virus, and other respiratory viruses: a systematic review of diagnostic accuracy and clinical impact studies. Clin Infect Dis. 2019;69(7):1243-1253.
- Lewinski MA, Alby K, Babady NE, et al. Exploring the utility of multiplex infectious disease panel testing for diagnosis of infection in different body sites: a joint report of the Association for Molecular Pathology, American Society for Microbiology, Infectious Diseases Society of America, and Pan American Society for Clinical Virology. J Mol Diagn. 2023;25(12):857-875.
- Liu W, Lee L. Toward rapid and accurate molecular diagnostics at home. Adv Mater. 2023;35:2206525.
Copyright © 2024 McMahon Publishing, 545 West 45th Street, New York, NY 10036. Printed in the USA. All rights reserved, including the right of reproduction, in whole or in part, in any form.

Download to read this article in PDF document:![]() Rapid Molecular Diagnostics for Respiratory Tract Infections: A Conceptual and Technical Overview
Rapid Molecular Diagnostics for Respiratory Tract Infections: A Conceptual and Technical Overview