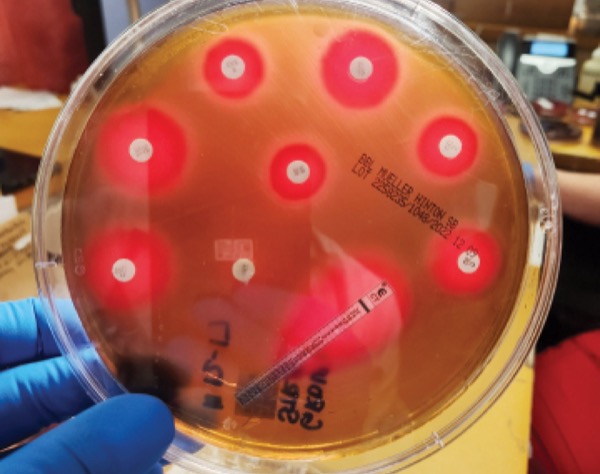
Antimicrobial resistance is part of natural selection and poses a serious problem in clinical medicine. Traditional methods for identifying microorganisms and testing for resistance to antimicrobials involve obtaining specimens (eg, blood, tissue, urine, pus, sputum) for culture followed by growing the organisms with antimicrobial agents to determine susceptibilities to antimicrobial agents. Although this process is very effective, it frequently takes 3 to 4 days before the clinician will have results allowing streamlining of antibiotics from broad- to narrow-spectrum agents.
Over the past decade, the field of rapid diagnostic testing has transformed the practice of clinical microbiology and medicine. Identification and susceptibility of bacteria and yeasts in blood is now possible within 1 to 2 days using rapid molecular techniques.1,2
This article focuses on bloodstream infections commonly seen in the hospital setting and discusses newer diagnostic technology that the clinician can use for both diagnosis and rapid determination of antibiotic resistance. Drug-resistant mycobacterial (tuberculosis), parasitic, or fungal infections (other than Candida spp) will not be discussed. The article also discusses newer antibiotics and uses of older antibiotics for treatment of multidrug-resistant (MDR) organisms.
Rapid Diagnostic and Antibiotic Susceptibility Tests
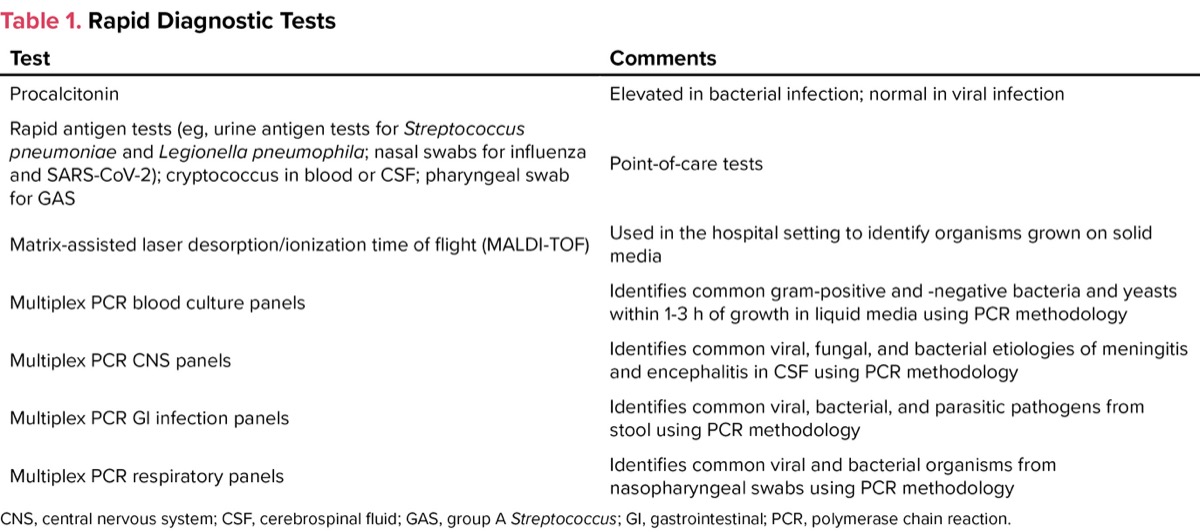
What is a rapid diagnostic test (RDT) in infectious diseases terminology? One definition is that it provides results to the clinician within 4 to 6 hours. Using this definition, there are several tests that qualify (Table 1). When evaluating a patient with an acute infection, it is sometimes difficult to distinguish a viral infection from a bacterial one. Use of a biomarker such as procalcitonin (Table 1) may be helpful in these situations because procalcitonin frequently is elevated in bacterial infections but not in viral ones.3 RDTs are used to identify specific organisms, whether they are viral, bacterial, fungal, or parasitic. These tests are designed to detect antigens, antibodies, or nucleic acid sequences that are specific to the organism in question.
| Table 1. Rapid Diagnostic Tests | |
| Test | Comments |
|---|---|
| Procalcitonin | Elevated in bacterial infection; normal in viral infection |
| Rapid antigen tests (eg, urine antigen tests for Streptococcus pneumoniae and Legionella pneumophila; nasal swabs for influenza and SARS-CoV-2); cryptococcus in blood or CSF; pharyngeal swab for GAS | Point-of-care tests |
| Matrix-assisted laser desorption/ionization time of flight (MALDI-TOF) | Used in the hospital setting to identify organisms grown on solid media |
| Multiplex PCR blood culture panels | Identifies common gram-positive and -negative bacteria and yeasts within 1-3 h of growth in liquid media using PCR methodology |
| Multiplex PCR CNS panels | Identifies common viral, fungal, and bacterial etiologies of meningitis and encephalitis in CSF using PCR methodology |
| Multiplex PCR GI infection panels | Identifies common viral, bacterial, and parasitic pathogens from stool using PCR methodology |
| Multiplex PCR respiratory panels | Identifies common viral and bacterial organisms from nasopharyngeal swabs using PCR methodology |
| CNS, central nervous system; CSF, cerebrospinal fluid; GAS, group A Streptococcus; GI, gastrointestinal; PCR, polymerase chain reaction. | |
Blood cultures have variable sensitivity depending on the diagnosis. For example, blood cultures are positive in roughly 25% to 40% of patients with sepsis, whereas they are rarely positive in patients with cellulitis. Positive blood cultures should help tailor antimicrobial therapy even before susceptibility testing is available. For example, broad-spectrum antibiotics such as vancomycin and piperacillin-tazobactam (or cefepime) commonly are started empirically for a patient with sepsis. If blood cultures obtained prior to starting antibiotics grow a gram-negative rod, then vancomycin can usually be discontinued (unless there are extenuating circumstances). Use of RDTs in blood cultures allows identification within 1 hour of growth in liquid media (after Gram stain has been done). Organisms identified depend on the vendor. Tables 2 through 4 list the common gram-positive and gram-negative bacteria and yeasts identified by this method.
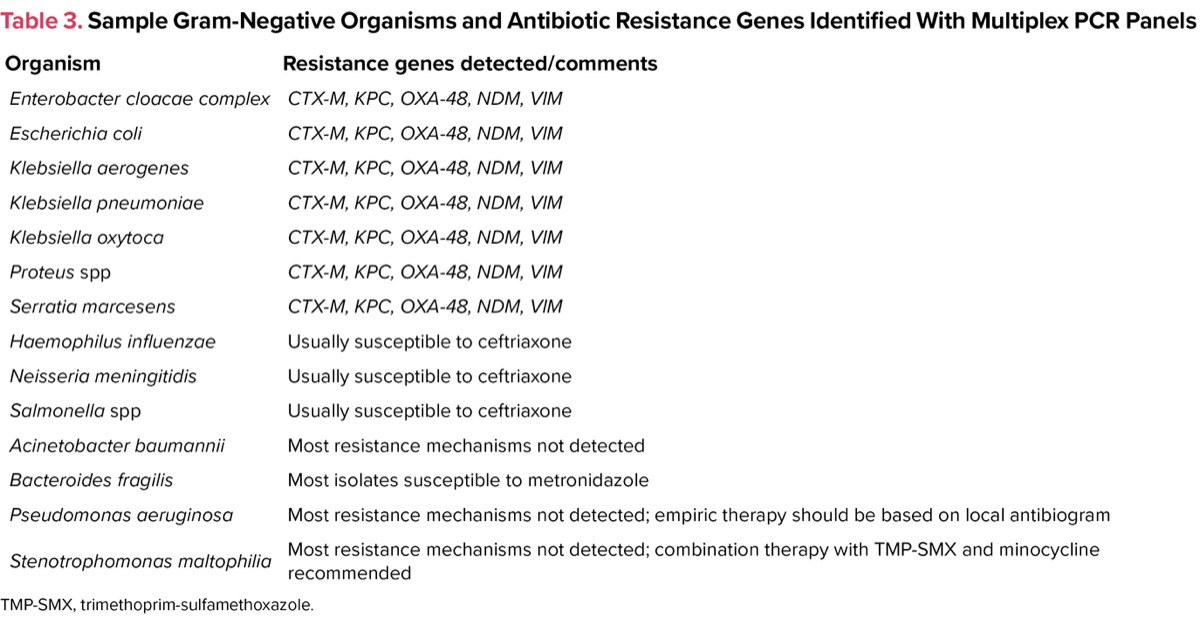
In addition to providing early identification of common organisms, multiplex polymerase chain reaction (PCR) can also give information regarding the presence or absence of common resistance genes. For example, specific genetic primers can determine the presence or absence of the mecA gene, which is the principal mechanism for resistance to methicillin and all other beta-lactam drugs (except ceftaroline and ceftobiprole). Blood cultures that grow gram-positive cocci would be probed with primers that detect the common gram-positive organisms (Table 2). For blood cultures growing gram-negative rods, these cultures are probed with a different set of primers that are specific for the common gram-negative organisms and many of the common resistance mechanisms (Table 3).
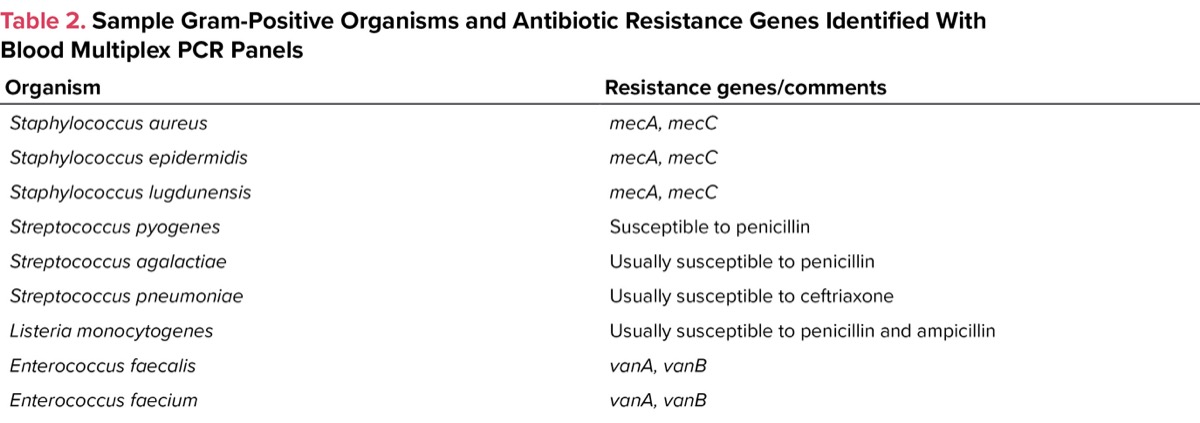
| Table 2. Sample Gram-Positive Organisms and Antibiotic Resistance Genes Identified With Blood Multiplex PCR Panels | |
| Organism | Resistance genes/comments |
|---|---|
| Staphylococcus aureus | mecA, mecC |
| Staphylococcus epidermidis | mecA, mecC |
| Staphylococcus lugdunensis | mecA, mecC |
| Streptococcus pyogenes | Susceptible to penicillin |
| Streptococcus agalactiae | Usually susceptible to penicillin |
| Streptococcus pneumoniae | Usually susceptible to ceftriaxone |
| Listeria monocytogenes | Usually susceptible to penicillin and ampicillin |
| Enterococcus faecalis | vanA, vanB |
| Enterococcus faecium | vanA, vanB |
| Table 3. Sample Gram-Negative Organisms and Antibiotic Resistance Genes Identified With Multiplex PCR Panels | |
| Organism | Resistance genes detected/comments |
|---|---|
| Enterobacter cloacae complex | CTX-M, KPC, OXA-48, NDM, VIM |
| Escherichia coli | CTX-M, KPC, OXA-48, NDM, VIM |
| Klebsiella aerogenes | CTX-M, KPC, OXA-48, NDM, VIM |
| Klebsiella pneumoniae | CTX-M, KPC, OXA-48, NDM, VIM |
| Klebsiella oxytoca | CTX-M, KPC, OXA-48, NDM, VIM |
| Proteus spp | CTX-M, KPC, OXA-48, NDM, VIM |
| Serratia marcesens | CTX-M, KPC, OXA-48, NDM, VIM |
| Haemophilus influenzae | Usually susceptible to ceftriaxone |
| Neisseria meningitidis | Usually susceptible to ceftriaxone |
| Salmonella spp | Usually susceptible to ceftriaxone |
| Acinetobacter baumannii | Most resistance mechanisms not detected |
| Bacteroides fragilis | Most isolates susceptible to metronidazole |
| Pseudomonas aeruginosa | Most resistance mechanisms not detected; empiric therapy should be based on local antibiogram |
| Stenotrophomonas maltophilia | Most resistance mechanisms not detected; combination therapy with TMP-SMX and minocycline recommended |
| TMP-SMX, trimethoprim-sulfamethoxazole. | |
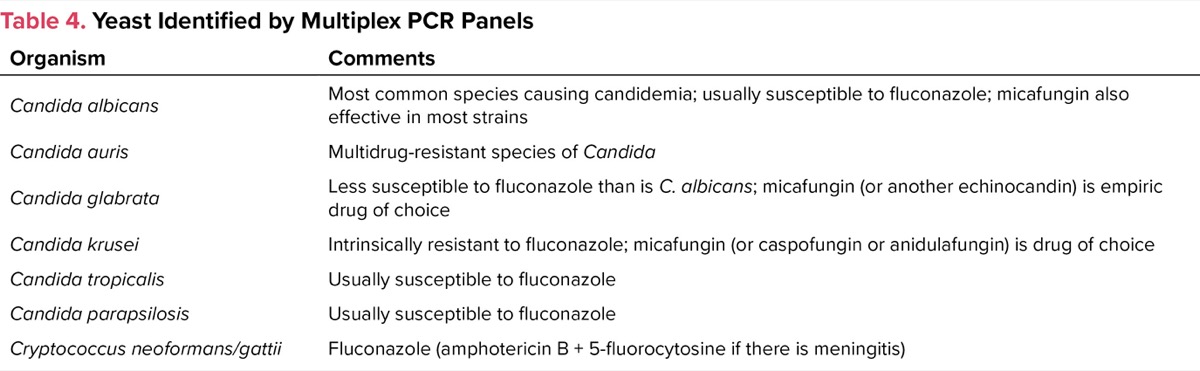
| Table 4. Yeast Identified by Multiplex PCR Panels | |
| Organism | Comments |
|---|---|
| Candida albicans | Most common species causing candidemia; usually susceptible to fluconazole; micafungin also effective in most strains |
| Candida auris | Multidrug-resistant species of Candida |
| Candida glabrata | Less susceptible to fluconazole than is C. albicans; micafungin (or another echinocandin) is empiric drug of choice |
| Candida krusei | Intrinsically resistant to fluconazole; micafungin (or caspofungin or anidulafungin) is drug of choice |
| Candida tropicalis | Usually susceptible to fluconazole |
| Candida parapsilosis | Usually susceptible to fluconazole |
| Cryptococcus neoformans/gattii | Fluconazole (amphotericin B + 5-fluorocytosine if there is meningitis) |
Common Bacteria and Fungi Causing Bloodstream Infections
The ESKAPE organisms Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter spp are encountered commonly in clinical practice (Table 5).4 MDR is common in all these organisms. Fortunately, all the ESKAPE organisms are detected by most of the commercially available multiplex PCR platforms for positive blood cultures. As such, if there is major antibiotic resistance, it will be detected quickly, and timely changes to treatment can usually be made. However, if the culture is not from blood, then it is likely that identification and susceptibility testing will be done by conventional methods, which are much slower.
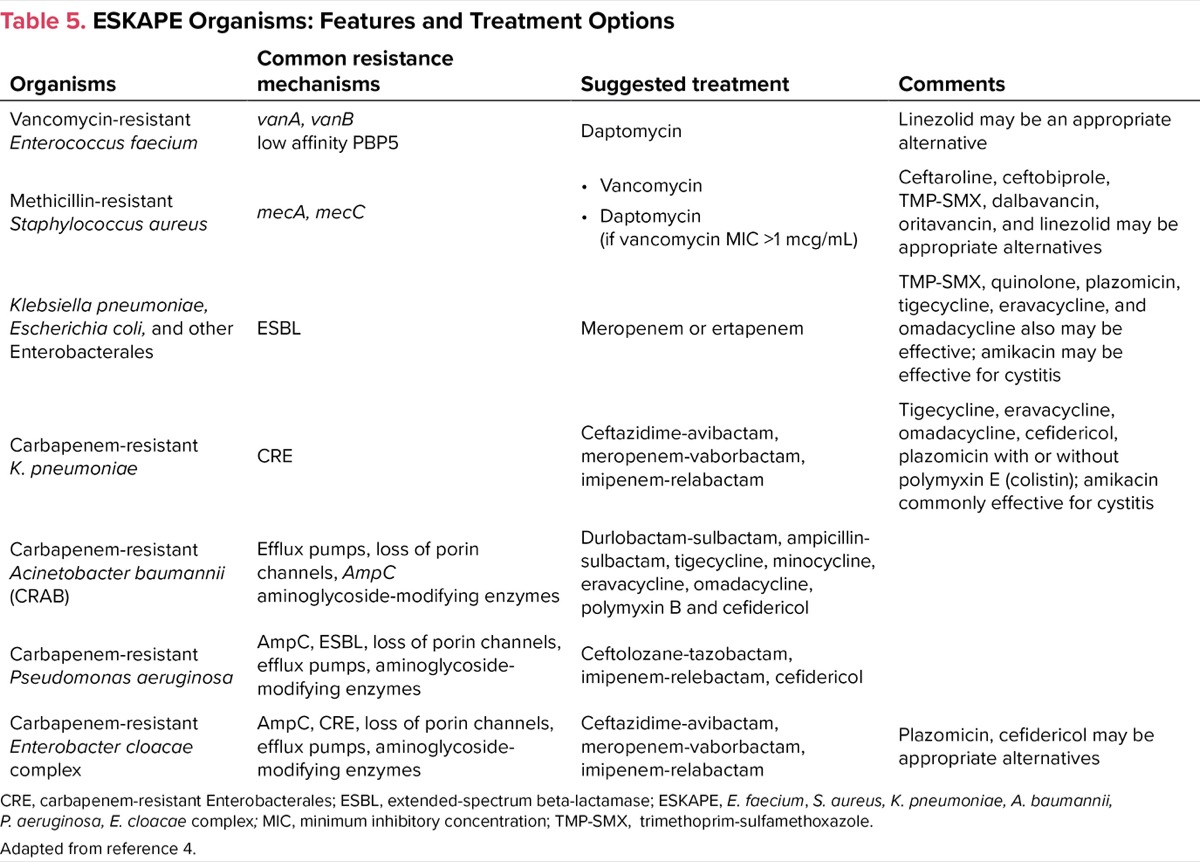
| Table 5. ESKAPE Organisms: Features and Treatment Options | |||
| Organisms | Common resistance mechanisms | Suggested treatment | Comments |
|---|---|---|---|
| Vancomycin-resistant Enterococcus faecium | vanA, vanB low affinity PBP5 | Daptomycin | Linezolid may be an appropriate alternative |
| Methicillin-resistant Staphylococcus aureus | mecA, mecC |
| Ceftaroline, ceftobiprole, TMP-SMX, dalbavancin, oritavancin, and linezolid may be appropriate alternatives |
| Klebsiella pneumoniae, Escherichia coli, and other Enterobacterales | ESBL | Meropenem or ertapenem | TMP-SMX, quinolone, plazomicin, tigecycline, eravacycline, and omadacycline also may be effective; amikacin may be effective for cystitis |
| Carbapenem-resistant K. pneumoniae | CRE | Ceftazidime-avibactam, meropenem-vaborbactam, imipenem-relabactam | Tigecycline, eravacycline, omadacycline, cefidericol, plazomicin with or without polymyxin E (colistin); amikacin commonly effective for cystitis |
| Carbapenem-resistant Acinetobacter baumannii (CRAB) | Efflux pumps, loss of porin channels, AmpC aminoglycoside-modifying enzymes | Durlobactam-sulbactam, ampicillin-sulbactam, tigecycline, minocycline, eravacycline, omadacycline, polymyxin B and cefidericol | |
| Carbapenem-resistant Pseudomonas aeruginosa | AmpC, ESBL, loss of porin channels, efflux pumps, aminoglycoside-modifying enzymes | Ceftolozane-tazobactam, imipenem-relebactam, cefidericol | |
| Carbapenem-resistant Enterobacter cloacae complex | AmpC, CRE, loss of porin channels, efflux pumps, aminoglycoside- modifying enzymes | Ceftazidime-avibactam, meropenem-vaborbactam, imipenem-relabactam | Plazomicin, cefidericol may be appropriate alternatives |
| CRE, carbapenem-resistant Enterobacterales; ESBL, extended-spectrum beta-lactamase; ESKAPE, E. faecium, S. aureus, K. pneumoniae, A. baumannii, P. aeruginosa, E. cloacae complex; MIC, minimum inhibitory concentration; Adapted from reference 4. | |||
Risk factors for vancomycin-resistant enterococci (VRE) infections include malignancy, organ transplantation, gastrointestinal surgeries, broad-spectrum antimicrobials, prolonged hospitalization, and admission to a long-term care facility. Treatment options for serious infections are limited (Table 5). Methicillin-resistant S. aureus (MRSA) infections are common in both the community and hospital settings. Vancomycin resistance with MRSA is uncommon, but treatment failure with MRSA isolates having a minimum inhibitory concentration (MIC) greater than 1 mcg/mL is common. Daptomycin, ceftaroline, and telavancin (Vibativ, Cumberland Pharmaceuticals) are reasonable options in these cases. Dalbavancin (Dalvance, AbbVie) and oritavancin (Orbactiv, Melinta Therapeutics) are considerations for outpatient management. Ceftobiprole (Zevtera, Basilea Pharmaceutica International) was approved recently for the treatment of S. aureus bloodstream infections as well as for infections involving the skin and respiratory tract (pneumonia). Like ceftaroline, ceftobiprole has activity against both methicillin-susceptible S. aureus and MRSA.
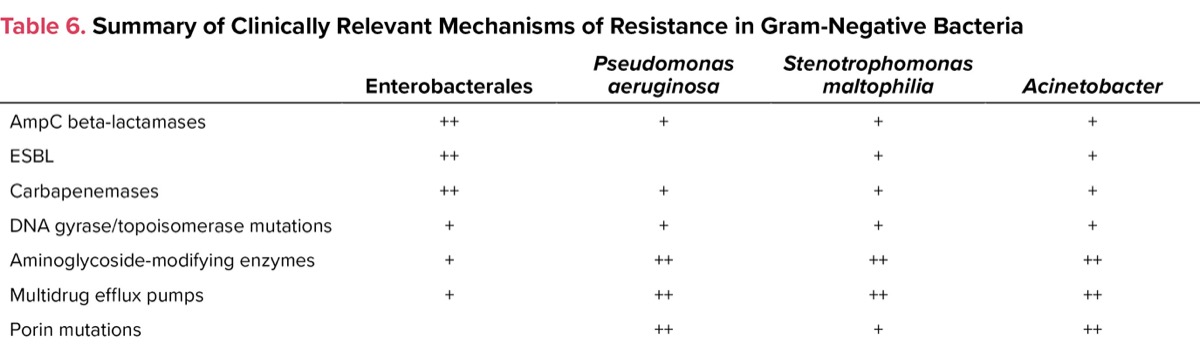
Resistance in gram-negative organisms is a more complicated process than with gram-positive bacteria. In general, resistance may be related to mutations that affect the entry of antibiotics into bacteria (ie, porins); altered binding to receptor molecules; enzymatic degradation (eg, beta-lactamases); or increased activity of efflux pumps that remove the antibiotic from the bacteria (Table 6).4-5 Much attention has been given to beta-lactamases, which are enzymes that hydrolyze the beta-lactam ring of penicillins, cephalosporins, and carbapenems, thereby rendering the drugs inactive. The clinically important beta-lactamases in gram-negative organisms include AmpC, extended-spectrum beta-lactamases (ESBL), and carbapenemases. Enterobacter spp, Serratia marcesens, and Citrobacter spp commonly possess AmpC-type beta-lactamases. These organisms typically are resistant to first-, second-, and third-generation cephalosporins. The AmpC gene is chromosomal in Enterobacter, Serratia, and Citrobacter and is inducible by beta-lactams. AmpC is either poorly expressed or not expressed in Escherichia coli and Klebsiella spp, respectively, but the gene can be acquired from a plasmid of another organism, thus giving it the same phenotype as Enterobacter. ESBL-producing bacteria can hydrolyze penicillins, cephalosporins, and monobactams. Fortunately, the carbapenems and cefepime are effective against AmpC-producing organisms, but derepression of AmpC with other resistance mechanisms can complicate the situation. ESBL-producing organisms often carry various genes on plasmids that make them resistant to other classes of antibiotics, such as aminoglycosides and fluoroquinolones. Although not part of the ESKAPE organisms, Stenotrophomonas maltophilia is increasingly a difficult pathogen to treat owing to its intrinsic resistance to most antibiotics (Table 6).4-6
| Table 6. Summary of Clinically Relevant Mechanisms of Resistance in | ||||
| Enterobacterales | Pseudomonas aeruginosa | Stenotrophomonas maltophilia | Acinetobacter | |
|---|---|---|---|---|
| AmpC beta-lactamases | ++ | + | + | + |
| ESBL | ++ | + | + | |
| Carbapenemases | ++ | + | + | + |
| DNA gyrase/topoisomerase mutations | + | + | + | + |
| Aminoglycoside-modifying enzymes | + | ++ | ++ | ++ |
| Multidrug efflux pumps | + | ++ | ++ | ++ |
| Porin mutations | ++ | + | ++ | |
Carbapenemase-producing organisms are being recognized increasingly in Enterobacterales, most commonly K. pneumoniae, and are designated carbapenem-resistant Enterobacterales (CRE). The carbapenemases are a diverse group of beta-lactamases; the common types are designated serine carbapenemases (classes A, C, and D) and metallo–beta-lactamases (class B) because of the critical importance of the zinc ion to beta-lactamase activity. The latter class includes the New Delhi metallo–beta-lactamases. Because carbapenemase genes are located on plasmids, there is the potential for rapid transfer of the genetic material to other organisms. Also, cotransfer of other resistance genes from the same plasmid is common, making these organisms resistant to all beta-lactams, aminoglycosides, fluoroquinolones, and tetracyclines.
Ceftazidime-avibactam (Avycaz, AbbVie) has potent in vitro activity against most strains of CRE, and ESBL- and AmpC-producing Enterobacteriaceae. Ceftazidime-avibactam is active against class A, C, and D carbapenemases; it has no activity against the class B metallo–beta-lactamases (eg, New Delhi metallo–beta-lactamases). Meropenem-vaborbactam (Vabomere, Melinta Therapeutics) and imipenem-relebactam-cilastatin (Recarbrio, Merck) have excellent activity against most CRE, and ESBL- and AmpC-producing Enterobacterales. Like ceftazidime-avibactam, meropenem-vaborbactam, and imipenem-relebactam have poor activity against class B metallo–beta-lactamases. Fortunately, these organisms are uncommon in the United States but can be seen in people who have traveled to areas where bacterial resistance is more common. Plazomicin (Zemdri, Cipla Therapeutics) is an aminoglycoside that is active against most ESBL- and CRE-producing Enterobacterales and is stable against most aminoglycoside-modifying enzymes. It has activity against most class B metallo–beta-lactamases. It has been studied mainly in urinary tract infections UTIs), so there are limited efficacy and safety data for other types of infections. One advantage of this drug over the beta-lactam/beta-lactamase inhibitor combinations is once-daily dosing.
Efflux pumps are present in many bacteria and are responsible for removal of a variety of substances, including certain antimicrobial agents, from the cell. Upregulation of efflux pumps is a common mechanism of resistance for both P. aeruginosa and A. baumannii to carbapenems, particularly meropenem and doripenem, and to fluoroquinolones. Other forms of resistance seen with P. aeruginosa and Acinetobacter include aminoglycoside-modifying enzymes, and AmpC and metallo–beta-lactamases (Table 6).
Porins are outer membrane protein channels that allow certain substances, including some antibiotics, to penetrate the bacterial cell membrane and wall. Downregulation of the outer membrane proteins results in resistance of the organism to the drug (eg, imipenem-resistant P. aeruginosa). It is common for phenotypic resistance to certain antibiotics to be due to a variety of mechanisms, including carbapenemase or derepressed AmpC activity, in the presence of reduced expression of porins and upregulation of efflux pumps (Table 6).
Ceftolozane-tazobactam (Zerbaxa, Merck) has excellent activity against P. aeruginosa, including many strains that are resistant to carbapenems, cefepime, and piperacillin-tazobactam. The addition of tazobactam provides enhanced activity against many MDR E. coli, but the combination drug has poor activity against ESBL-producing Enterobacterales and no activity against CRE organisms. Plazomicin and cefidericol (Fetroja, Shionogi) have good activity against MDR P. aeruginosa.
Infections with A. baumannii tend to be resistant to multiple agents. Treatment of carbapenem-resistant A. baumannii (CRAB) is difficult.4,5 The common mechanisms of resistance involve a combination of decreased porin channels, AmpC beta-lactamases, carbapenemases, and increased efflux pumps (Table 6). Therefore, drugs such as meropenem, ceftazidime-avibactam, ceftolozane-tazobactam, imipenem-relebactam, and meropenem-vaborbactam frequently are ineffective. Aminoglycosides, including plazomicin, are not active against CRAB. High-dose ampicillin-sulbactam has been the drug of choice for CRAB, but that recommendation may be changing. Sulbactam is the active component. It works by binding to penicillin-binding proteins (PBPs) PBP1 and PBP3. Unfortunately, sulbactam is hydrolyzed by a variety of beta-lactamases present in CRAB. The newly approved antibiotic durlobactam-sulbactam (Xacduro, Innovia Specialty Therapeutics) is a beta-lactam/beta-lactamase inhibitor combination with activity against CRAB, including most isolates that are resistant to sulbactam, minocycline, and colistin.5 Durlobactam is a potent inhibitor of the beta-lactamases associated with CRAB, thus allowing sulbactam to avoid hydrolysis and bind to PBP1 and PBP3. Alternative treatments for this difficult-to-treat organism include a combination of polymyxin with one of the tetracyclines (minocycline, tigecycline, eravacycline [Xerava, Tetraphase], or omadacycline [Nuzyra, Paratek]). Cefidericol, a sideraphore cephalosporin, has good activity against MDR A. baumannii and MDR P. aeruginosa. The clinical data of cefidericol in patients with CRAB infections are limited. This drug is also stable against AmpC-, ESBL-, and carbapenemase–beta-lactamases.
The RDTs for gram-negative organisms are most helpful for the Enterobacterales because resistance in this group of organisms typically is mediated via beta-lactamases (Table 6). These tests are less useful in the management of infections (with bacteremia) caused by A. baumannii, S. maltophilia, and P. aeruginosa since resistance is mediated commonly by other mechanisms, including porin channels and efflux pumps (Table 6). The RDTs currently available do not detect the genes that are involved in these resistance processes.
Bloodstream infections with yeasts, usually Candida spp, are becoming much more common in the hospital and community settings, particularly as central catheter use in the outpatient setting has increased. Patients receiving broad-spectrum antibiotics, steroids, and total parenteral nutrition are at particularly high risk. Cancer patients with chemotherapy-associated neutropenia are also at high risk. Multiplex PCR has facilitated rapid diagnosis of candidemia to the species level (Table 4). This information facilitates empiric treatment while awaiting formal susceptibility testing, which is usually done at a reference laboratory and has a turnaround time of several days.
Older antibiotics have a role in the treatment of MDR infections involving the lower urinary tract (i.e., cystitis). Nitrofurantoin is active against most E. coli, which is the most common organism associated with UTIs. The drug is more active against ESBL-producing E. coli than CRE E. coli. Nitrofurantoin is less active against both ESBL-producing and CRE K. pneumoniae. Fosfomycin is active against both ESBL-producing and CRE E. coli. It is less active against ESBL-producing and CRE K. pneumoniae, but the drug has better activity than nitrofurantoin against these organisms. Nitrofurantoin and fosfomycin should not be used in patients with pyelonephritis or bacteremia. Nitrofurantoin is less effective in the presence of a glomerular filtration rate less than 40 mL per minute. Amikacin is frequently active against ESBL-producing E. coli and K. pneumoniae and can be given as a single dose (15 mg/kg of body weight) for treatment of uncomplicated cystitis.
Conclusion
Appropriate use of RDTs should help reduce the inappropriate use of broad-spectrum antimicrobial agents, leading to improvements in antimicrobial susceptibilities. The use of multiplex PCR for identification of common bloodstream infections has greatly improved the care of patients with sepsis by allowing timely deescalation of broad-spectrum antibiotics for some, and triggering timely use of newer antibiotics in certain patients with MDR organisms. A few new antibiotics have been approved recently, and more drugs should become available in the near future. Older drugs, including nitrofurantoin, fosfomycin, and amikacin, are effective against many ESBL-producing and some CRE E. coli that cause lower UTIs.
References
- Timbrook TT, Morton JB, McConeghy KW, et al. The effect of molecular rapid diagnostic testing on clinical outcomes in bloodstream infections: a systematic review and meta-analysis. Clin Infect Dis. 2017;64(1):15-23.
- van Belkum A, Burnham CAD, Rossen JWA. et al. Innovative and rapid antimicrobial susceptibility testing systems. Nat Rev Microbiol. 2020;18:299-311.
- Gregoriano C, Heilmann E, Molitor A, et al. Role of procalcitonin use in the management of sepsis. J Thorac Dis. 2020;12 (suppl 1):S5-S15.
- Miller WR, Arias CA. ESKAPE pathogens: antimicrobial resistance, epidemiology, clinical impact and therapeutics. Nat Rev Microbiol. 2024 Jun 3. https://doi.org/10.1038/s41579-024-01054-w
- Tamma PD, Aitken SL, Bonomo RA, et al. Infectious Diseases Society of America 2023 guidance on the treatment of antimicrobial resistant gram-negative infections. Clin Infect Dis. 2023:ciad428.
- Brooke JS. Advances in the microbiology of Stenotrophomonas maltophilia. Clin Microbiol Rev. 2021;34:10.
Copyright © 2024 McMahon Publishing, 545 West 45th Street, New York, NY 10036. Printed in the USA. All rights reserved, including the right of reproduction, in whole or in part, in any form.

Download to read this article in PDF document:![]() The Role of Rapid Diagnostic Testing and New Antimicrobials in Clinical Practice; Fighting Antimicrobial Resistance
The Role of Rapid Diagnostic Testing and New Antimicrobials in Clinical Practice; Fighting Antimicrobial Resistance